HDF5 format
HDF5 data format
The Eiger detector records images in the HDF5 data format. Instead of individual image files, the data is stored in container files of 100 images each:
-rw-r--r-- 1 opid30 jsbg 87M Feb 4 11:45 insulin_w1_5_1_master.h5 -rw-r--r-- 1 opid30 jsbg 304M Feb 4 11:45 insulin_w1_5_1_data_000001.h5 -rw-r--r-- 1 opid30 jsbg 313M Feb 4 11:45 insulin_w1_5_1_data_000002.h5 -rw-r--r-- 1 opid30 jsbg 328M Feb 4 11:45 insulin_w1_5_1_data_000003.h5 -rw-r--r-- 1 opid30 jsbg 350M Feb 4 11:45 insulin_w1_5_1_data_000004.h5 -rw-r--r-- 1 opid30 jsbg 367M Feb 4 11:45 insulin_w1_5_1_data_000005.h5
Viewing
For viewing, use either albula or adxv.
Note that with albula you need to open the *_master.h5 file, while with adxv you need to open one of the individual *_data_00000X.h5 files.
To automatically display images as they collected ("follow mode"), use albula and enable the "Eiger Monitor" in the "Auto Load" menu by clicking on it (such that it's printed in bold).
Note that these images don't have resolution information in the tooltip. Load the actual image files to see the resolution.
And make sure you use the "Auto Contrast" button in the "Tools" menu on the right.
Adxv can display the images as well, but only if launched via the Start menu, or by typing adxv_sc3 instead of adxv on the command line (because it's a special version of adxv that is running on the sample changer PC).
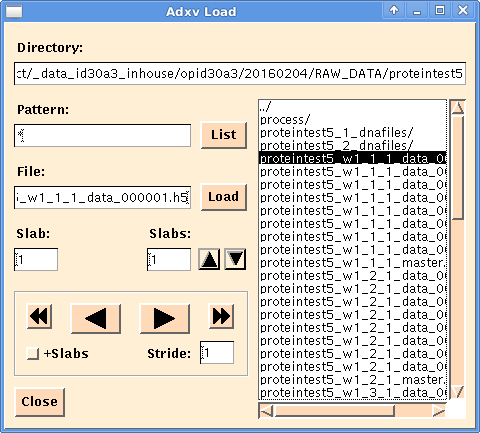
Once a *_data_00000x.h5 is opened in adxv, additional buttons appear in the "Load" dialog: If the "+Slabs" button is not actived, the left/right arrow switch between files (as with other image formats), but when it is actived, the left/right arrows switch between images within a hdf5 file.
Additionally, the "Slabs:" field allows to automatically sum up images for easier evaluation of the diffraction quality: e.g., the summation of 10 images of 0.1° oscillation each results in a displayed image that corresponds to 1° oscillation.
Processing
For processing with XDS, it is necessary to install the H5ToXds tool from Dectris (see here for more details). Then it is sufficient to point XDS to the HDF5 master file by using the following line in XDS.INP, and the conversion to CBF files will happen automatically in the background:
NAME_TEMPLATE_OF_DATA_FRAMES= insulin_w1_5_1_??????.h5 ! HDF5
Note that this string matches only the *_master.h5 file, but nevertheless XDS insists in having wildcards in NAME_TEMPLATE_OF_DATA_FRAMES.
If you want to use mosflm, there is a converter to CBF files for Linux and OSX available for download on the mosflm page, which unlike H5toXDS produces CBF files with miniCBF headers:
http://www.mrc-lmb.cam.ac.uk/harry/imosflm/ver721/downloads/eiger2cbf-linux.zip
http://www.mrc-lmb.cam.ac.uk/harry/imosflm/ver721/downloads/eiger2cbf-linux-old.zip
http://www.mrc-lmb.cam.ac.uk/harry/imosflm/ver721/downloads/eiger2cbf-osx.zip
To process HDF5 files with autoPROC, use this command:
process -h5 samplename_master.h5



