- Home
- News
- Spotlight on Science
- How an icosahedral...
How an icosahedral virus packages its RNA genome
20-12-2018
The spontaneous reconstitution of a plant icosahedral virus was probed by time-resolved small-angle X-ray scattering. The RNA genome acts as an assembly template for the protein subunits making up the viral capsid. A detailed understanding of the molecular mechanisms of viral self-assembly pathways could lead to the development of new therapeutic strategies against viral replication and to the design of new nanocontainers for drug delivery.
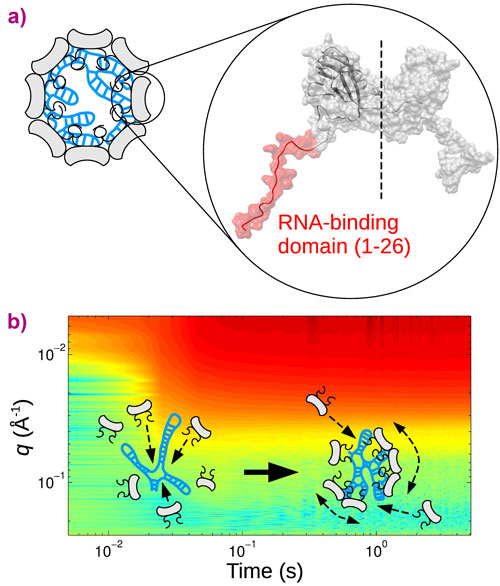
Cowpea chlorotic mottle virus (CCMV) is an icosahedral single-stranded RNA virus infecting a variety of beans. Four segments of RNA encoding the viral genome are shared between three distinct protein shells, called capsids, necessary to the viral replication. The capsids, which protect the RNA genome, are made up of 90 dimeric subunits (Figure 1a) arrayed in a structure with 20 faces, an icosahedron. Each subunit possesses two flexible, highly positively charged domains that interact electrostatically with the genome. A delicate balance of interactions between subunits, combining a short-range mostly hydrophobic attraction and a long-range electrostatic repulsion, holds the capsid together. The survival of the virus partly relies on its ability to self-assemble rapidly, in an error-free manner, inside the host cell. Although coarse-grained simulations have identified different pathways leading to the assembled virus from its components, very few experimental measurements were available up to now, due to the difficulty of following the fate of biological molecules over a wide range of timescales.
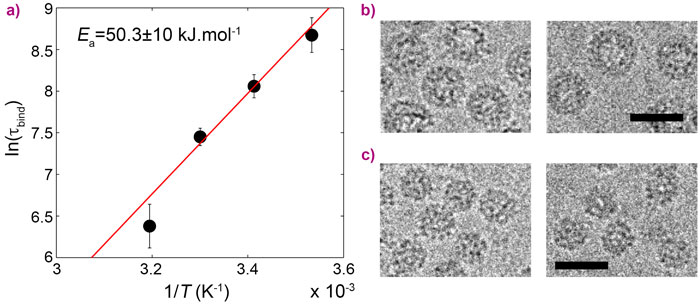
In this work, CCMV was reconstituted from purified protein subunits and genomic RNA, and its spontaneous assembly was followed by time-resolved small-angle X-ray scattering (SAXS) at beamline ID02. The forward scattering intensities allowed an estimation of the mean number of subunits bound to the genome as a function of time and, subsequently, the binding timescale, τbind, while structural information was inferred from the form factors. An empty capsid self-assembles sequentially and requires precise positioning of the subunits over the course of the assembly [1, 2]. For a full virus, the genome acts as an assembly template and captures a large number of subunits to form a disordered nucleoprotein complex (Figure 1b). The latter slowly self-organises, typically during one hour at room temperature, while continuing to capture the missing subunits, and eventually leads to a fully-assembled virus. Equilibrium measurements at various subunit-to-genome mass ratios revealed that the binding energy of subunits on the genome is moderate (i.e., ~7 kBT0 where kB is the Boltzmann constant and T0 the room temperature). In vivo, a weak binding energy may allow the subunits to select the viral genome specifically, which is a crucial feature in the crowded environment of the cytoplasm. Relaxation experiments carried out at different temperatures showed that the self-organisation of nucleoprotein complexes into viruses requires a high energy barrier to be crossed, Ea, estimated to be 20 kBT0 (Figure 2a). The assembled viruses (Figure 2b) are structurally indistinguishable from native viruses. In particular, they display a hollow core as the genome is packed against the interior wall of the capsid due to the electrostatic interactions. In the last series of experiments, a synthetic polyelectrolyte, namely poly(styrene sulfonic acid) (PSS), was packaged into capsids (Figure 2c) under conditions where viruses could not build up. The equilibrium state was reached in only two minutes. PSS has a hydrophobic backbone so its excluded volume is smaller than that of RNA. Its compaction therefore requires less energy and its templating effect is enhanced. PSS-filled capsids were fairly spherical and regular but exhibited defects that suggest the icosahedral symmetry is not respected.
The elucidated pathways of genome packaging might be advantageous to the virus’s survival for a twofold reason: (i) The building blocks are first associated rapidly due to nonspecific but moderate electrostatic interactions, and (ii) the high energy barrier between nucleoprotein complexes and viruses provides enough time to select the viral genome and correct the errors of assembly.
Principal publication and authors
Nonequilibrium self-assembly dynamics of icosahedral viral capsids packaging genome or polyelectrolyte, M. Chevreuil (a, b), D. Law-Hine (a), J. Chen (a), S. Bressanelli (b), S. Combet (c), D. Constantin (a), J. Degrouard (a), J. Möller (d), M. Zeghal (a) and G. Tresset (a), Nat. Commun. 9, 3071 (2018); doi: 10.1038/s41467-018-05426-8.
(a) Laboratoire de Physique des Solides, Orsay (France)
(b) Institute for Integrative Biology of the Cell (I2BC), Gif-sur-Yvette (France)
(c) Laboratoire Léon Brillouin (LLB), Gif-sur-Yvette (France)
(d) ESRF
References
[1] G. Tresset et al., J. Am. Chem. Soc. 135, 15373–15381 (2013).
[2] D. Law-Hine et al., Soft Matter 12, 6728–6736 (2016).