- Home
- News
- Spotlight on Science
- A new class of signalling...
A new class of signalling molecules revealed by structure solution exploiting the anomalous scattering of sulphur
18-04-2013
FAM3B PANDER is involved in the regulation of glucose homeostasis and β cell function and FAM3C ILEI is implicated in cancer. They had both been predicted to be classical cytokines with three dimensional structures based on a 4-helix bundle. The crystal structure of FAM3B PANDER, solved by exploiting the anomalous scattering of sulphur atoms inherent to the amino acid sequence of the protein has revealed that, contrary to expectations, PANDER adopts a novel, globular β-β-α fold that is conserved throughout the FAM3 superfamily. This superfamily most likely represents a novel class of signalling molecules with a mode of action distinct to that of the well-known four-helix bundle cytokines.
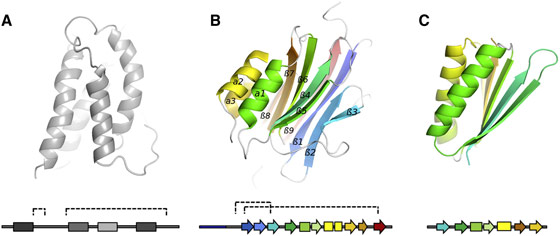
The FAM3 superfamily of signalling molecules comprises four currently identified members, FAM3A, -B, -C, and -D, all of which are predicted to adopt three-dimensional structures based on an up-up-down-down 4-helix bundle stabilised by two disulphide bridges ([1], Figure 1). Of the four FAM3 superfamily members, FAM3B and FAM3C have been studied extensively, have been renamed FAM3B PANDER (pancreatic derived factor) and FAM3C ILEI (interleukin-like EMT inducer) and are involved in glucose homeostasis and cancer, respectively. It has also been proposed that the presence of FAM3C ILEI could be exploited as a predictive marker for breast cancer [2]. Both FAM3B PANDER and FAM3C ILEI represent potential targets for drugs aimed at treating the diseases in which they are implicated.
Structural information is of fundamental importance both in the assignment of function (i.e. mode of action) of a biological macromolecule and in the design of molecules (i.e. drugs) to modify this. The crystal structure of FAM3B PANDER (Figure 1) was recently solved using single crystal diffraction data collected at beamline ID29 at a wavelength designed to exploit the small anomalous signal due to sulphur atoms inherent to the amino acid sequence of the protein in the structure solution process. Surprisingly, the crystal structure of FAM3B shows no relationship to the expected four-helix bundle fold and instead adopts a novel, globular β-β-α fold similar only to that of the theoretical, ab initio designed protein TOP7 [3] (Figure 1).
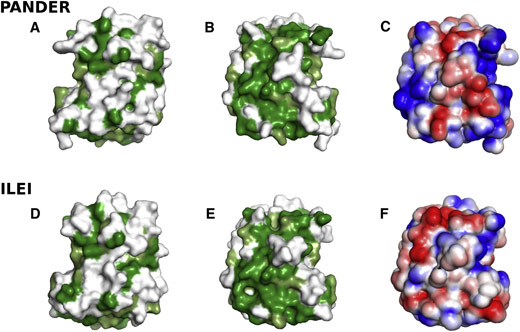
A structure-based amino acid sequence alignment of several FAM3 proteins revealed that the novel fold observed for FAM3B PANDER is likely to be conserved throughout superfamily members (i.e. FAM3A, -B, -C, and -D). Moreover, of two possible FAM3B PANDER active sites identified using computer prediction, one is a small, closed pocket lined by amino acid residues highly conserved in FAM3 members suggesting that this area may be important to FAM3 functionality. While amino acids found on the surface of the structure of FAM3B PANDER are not generally highly conserved throughout the FAM3 superfamily, a homology model of the structure of FAM3C ILEI based on an alignment of its amino acid sequence with that of FAM3B PANDER indicates that the two proteins share large regions of conserved surface amino acids although the electrostatic properties of the two surfaces are likely to be different (Figure 2). This suggests that they may interact with a similar overall category of binding partners but target different subtypes.
Currently, neither a receptor nor a definitive signalling pathway has been identified for either FAM3B PANDER or FAM3C ILEI. This makes it difficult to ascertain the modes of action and exact functions of the two proteins. The novel β-β-α fold observed for FAM3B PANDER and the fact that it appears to be conserved for all FAM3 superfamily members suggests that FAM3 cytokines represent a novel class of signalling molecules, and provides a new tool in further research aimed at elucidating the function of FAM3 proteins.
Principal publication and authors
FAM3B PANDER and FAM3C ILEI represent a distinct class of signaling molecules with a non-cytokine-like fold, P. Johansson (a), J. Bernström (b), T. Gorman (c), L. Öster (a), S. Bäckström (b), F. Schweikart (b), B. Xu (b), Y Xue (a), L. Holmberg Schiavone (b), Structure 21, 306-313 (2013).
(a) Structure and Biophysics, Discovery Sciences, AstraZeneca, Mölndal (Sweden)
(b) Reagents and Assay Development, Discovery Sciences, AstraZeneca, Mölndal (Sweden)
(c) High-Content Biology, Discovery Sciences, AstraZeneca, Macclesfield (UK)
References
[1] Y. Zhu et al., Genomics 80, 144–150 (2002).
[2] T. Waerner, M. Alacakaptan, I. Tamir, R. Oberauer, A. Gal, T. Brabletz, M. Schreiber, M. Jechlinger, H. Beug. Cancer Cell 10, 227–239 (2006).
[3] B. Kuhlman, G. Dantas, G.C. Ireton, G. Varani, B.L. Stoddard and D. Baker. Science 302, 1364–1368 (2003).
Article written by G. Leonard, ESRF.
Top image: Cartoon representation of the long wavelength S-SAD structure of FAM3B PANDER.