- Home
- News
- Spotlight on Science
- Prevention of GroEL...
Prevention of GroEL ring separation results in functionally defective symmetric GroEL:GroES2 complexes
17-04-2018
The chaperonin GroEL and its cofactor GroES form a nanocage which facilitates the correct folding of other proteins. The two rings of GroEL transiently separate in an ATP-dependent reaction. Structures of a GroEL mutant with covalently linked rings were solved at the ESRF and supported the conclusion that ring separation is part of the chaperonin cycle.
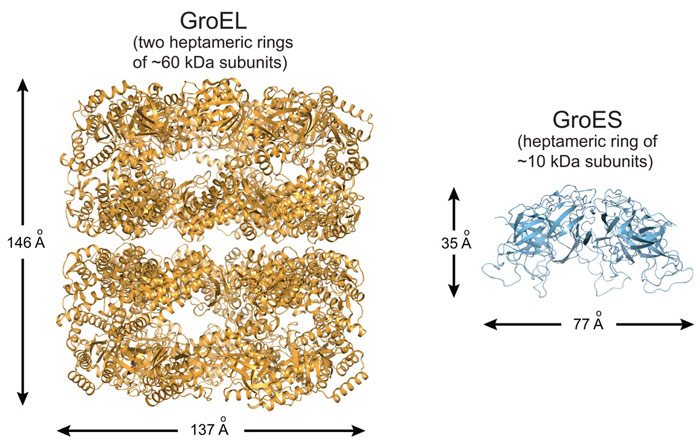
Proteins are synthesised as linear chains of amino acids but must fold into unique three-dimensional structures for functional activity. To fold efficiently on a biologically-relevant time scale, many proteins rely on assistance by molecular chaperones to avoid misfolding and aggregation in the crowded cellular environment. The bacterial chaperonin system, GroEL and its cofactor GroES, is arguably the best-studied molecular chaperone (Figure 1) [1]. GroEL is a cylindrical complex of ~800 kDa with ATPase activity, consisting of two heptameric rings stacked back-to-back. GroES, a single heptameric ring of ~10 kDa subunits, binds and unbinds from the ends of the GroEL cylinder in a reaction cycle allosterically controlled by the GroEL ATPase. A non-native substrate protein binds in the ring centre to hydrophobic residues exposed by the apical GroEL domains. Binding of GroES displaces the substrate protein into an enclosed cage for folding to occur unimpaired by aggregation. How ring opening and closing is orchestrated by ATP binding and hydrolysis in the two rings, and the exact nature of the structural intermediates in the folding cycle are only partially understood. Specifically, whether the two rings function sequentially in protein folding (asymmetric cycle, GroEL:GroES complexes populated) or in parallel (symmetric cycle, GroEL:GroES2 complexes populated) has been a matter of debate (Figure 2) [1].
 |
|
Figure 1. Ribbon representation of the GroEL tetradecamer complex (left panel; PDB 1XCK) and the GroES heptamer (right panel; PDB 4PKO). |
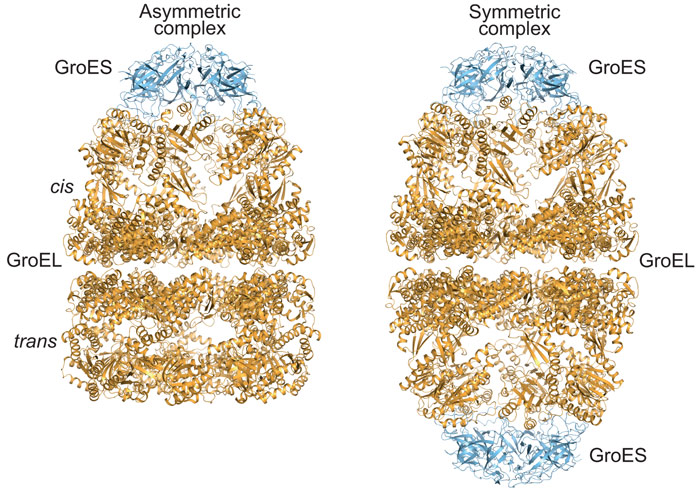
Our analysis of the GroEL/GroES reaction revealed that the two GroEL rings transiently separate during the chaperonin folding cycle. Ring separation occurs upon ATP binding to the trans-ring of the asymmetric GroEL:GroES/ADP complex (Figure 2) and results in ring exchange between GroEL complexes. To test the functional relevance of this step in the chaperonin cycle, we engineered cysteine residues into the GroEL ring-ring interface, which formed covalent disulphide bonds preventing ring separation. Crystal structures of this cysteine mutant were determined in two conformational states using diffraction data collected at ESRF. Although GroEL readily crystallises in a variety of conditions, diffraction quality varies dramatically between crystals. We therefore took advantage of fully automated data collection on beamline ID30A-1 (MASSIF-1) to systematically screen a large number of crystals and to rationally identify the best-diffracting volumes in large crystals by X-ray rastering, as implemented in the automated data collection workflow [2]. The crystallographic models suggest that formation of the disulphide bonds does not distort the structure compared to wild-type GroEL, consistent with our biochemical characterisation, which revealed normal ATP-regulated inter-ring allostery. However, in contrast to wild-type GroEL, the disulphide-bonded GroEL mutant populates symmetric GroEL:GroES2 complexes. Furthermore, the mutant was defective in substrate protein release and binding. Consistent with the essential function of GroEL/GroES in Escherichia coli, the mutant could not substitute for wild-type GroEL in vivo, especially under heat stress, when pre-existent proteins unfold and exert pressure on the chaperonin system.
 |
|
Figure 2. Crystal structures of asymmetric (left panel; PDB 1PCQ) and symmetric (right panel; PDB 4PKO) complexes of GroEL and GroES in ribbon representation. |
The present study on bacterial GroEL/GroES provides a mechanistic model for the chaperonin reaction. Homologs of GroEL occur in mitochondria (Hsp60) and in chloroplasts (Cpn60). Mutations in the human Hsp60 cause hereditary spastic paraplegia and chloroplasts Cpn60 is required for the folding of Rubisco, the abundant photosynthetic enzyme that catalyses the fixation of atmospheric carbon dioxide.
Principal publication and authors
GroEL Ring Separation and Exchange in the Chaperonin Reaction, X. Yan (a), Q. Shi (b), A. Bracher (a), G, Miličić (a), A.K. Singh (a), F.U. Hartl, Hayer-Hartl M, Cell 172, 605-617 (2018); doi: 10.1016/j.cell.2017.12.010.
(a) Max Planck Institute of Biochemistry, Martinsried (Germany)
(b) iHuman Institute, Shanghai (China)
References
[1] M. Hayer-Hartl, A. Bracher, and F.U. Hartl, Trends Biochem. Sci. 41, 62-76 (2016).
[2] M.W. Bowler, et al., J. Synchrotron Radiat. 22, 1540-1547 (2015).



