- Home
- News
- Spotlight on Science
- Rpf2-Rrs1 complex...
Rpf2-Rrs1 complex in the biogenesis of the ribosome
11-01-2016
The biogenesis of ribosomes is a highly complex and hierarchical process mediated by more than 200 biogenesis factors. An important early step is the incorporation of the ribonucleoprotein 5S RNP (5S rRNA, RpL5 and RpL11) guided by the heterodimeric Rpf2-Rrs1 complex. The de novo structure of this complex has now been solved. The structure revealed the unique complementation of the Rpf2 Brix-domain by Rrs1 to form a tandem array of anticodon binding domains (ABDs). Fitting of the structure into a cryo-EM density revealed its association with the 5S RNP, the pre-ribosome, and the biogenesis factor Rsa4 and explained its function in ribosome biogenesis.
Ribosomes are the main industrial zones within cells representing over 1/3 of their dry mass and being responsible for most of their power consumption. The construction of these megadalton factories is a hierarchical operating process barely understood in its complexity and aided in eukaryotes by about 200 biogenesis factors. The maturation of the large 60S subunit requires more than 70 factors clustering with different ribosome assembly intermediates and forming specific interactions with pre-rRNA and R-proteins of the ribosome. One such interaction network is formed during biogenesis of the ribosomal central protuberance (CP) including the 5S RNP [1]. The 5S RNP is composed of an RNA of about 120 nucleotides (5S rRNA) and two associated proteins (RpL5 and RpL11) [2]. The 5S RNP gets recruited to the pre-ribosome at an early stage and has to rearrange during later maturation stages of the pre-60S ribosomal subunit [3]. The two co-working assembly factors Rpf2 and Rrs1 were known to assist in 5S RNP biogenesis [4] but their role and location on the pre-ribosome remained enigmatic [3,5].
We thus set out to solve the X-ray structure of the Rpf2-Rrs1 complex with data collection performed at the ESRF. Having gone through quite extensive construct optimisation and species selection we managed to produce well diffracting native crystals for the Aspergillus nidulans proteins. Ribosome biogenesis is a highly competetive scientific field and we felt that we had to push this project forward quickly. Therefore, we sought support from the newly established MASSIF-1 beamline ID30A-1 [6,7]. We managed to fully-automatically collect 70 native datasets. As molecular replacement failed, we measured Se-Met derived crystals for phasing by single anomalous dispersion (SAD). MASSIF-1 is set at the wavelength of 0.96 Å close to the selenium absorption edge and we were able to collect another 40 datasets within one instantly allocated session and the structure was automatically solved 9 times at a resolution of 1.5 Å.
 |
|
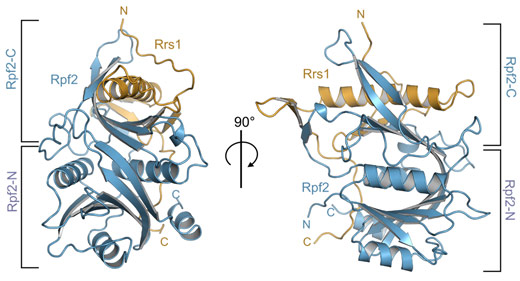
Figure 1. Overall structure of the Aspergillus nidulans Rpf2-Rrs1 complex. The two halves of Rpf2, Rpf2-N and Rpf2-C, are indicated. Rrs1 complements the fold of Rpf2-C. |
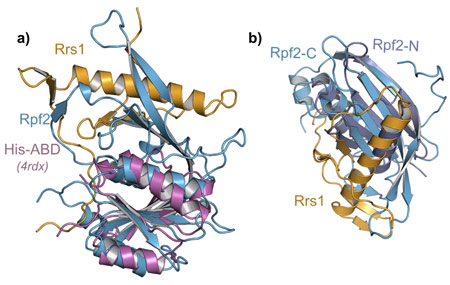
The structure of the Rpf2-Rrs1 complex revealed several unexpected features important for ribosome biogenesis (Figure 1). Rpf2 was known to have a so-called Brix-domain, but its intricate interaction interface and complementation by Rrs1 was only now unravelling. Brix-domains consist of two anticodon binding domains (ABDs) termed Rpf2-N and Rpf2-C (Figure 2a). However, Rpf2-C is an incomplete ABD and needs Rrs1 for domain complementation (Figure 2b).
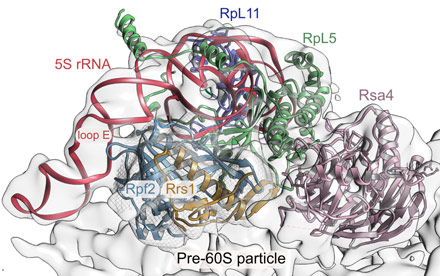
Recently, the cryo-EM structure of a pre-60S particle was solved and revealed an astonishing outward conformation of 5S RNP that is 180° rotated with respect to the mature conformation [3]. With the structure of the Rpf2-Rrs1 complex in hand, we could unambiguously place our high-resolution structure in unassigned density next to the 5S RNP (Figure 3). The Rpf2-Rrs1 complex is localised in close contact with the 5S rRNA, RpL5, and the ribosome biogenesis factor Rsa4. Fitting of Rpf2-Rrs1 revealed that Rpf2 can specifically read-out a conserved RNA-bulge (loop E) of 5S rRNA, supplying us with a first model for Brix-domain/RNA interaction. Further on, Rpf2-Rrs1 interacts with eukaryote specific insertions of RpL5 and with the so-called Ubl-domain of Rsa4. The intimate interaction of Rpf2-Rrs1 (and their elongated tails that could now be located on the pre-ribosome) with both 5S RNP and Rsa4 suggests the Rpf2-Rrs1 complex keeps the 5S RNP in its outward-conformation and that its removal is necessary to promote its rotation for pre-ribosome maturation.
Principal publication and authors
The structure of Rpf2-Rrs1 explains its role in ribosome biogenesis, S. Kharde, F.R. Calviño, A. Gumiero, K. Wild and I. Sinning, Nucleic Acids Res. 43, 7083–95 (2015); doi: 10.1093/nar/gkv640.
Heidelberg University Biochemistry Center (BZH), Heidelberg (Germany)
References
[1] F.R. Calviño et al., Nat. Commun. 6, 6510 (2015); doi: 10.1038/ncomms7510.
[2] A. Ben-Shem et al., Science 334, 1524–9 (2011); doi: 10.1126/science.1194294.
[3] C. Leidig et al., Nat. Commun. 5, 3491 (2014); doi: 10.1038/ncomms4491.
[4] J. Zhang et al., Genes Dev. 21, 2580–92 (2007); doi: 10.1101/gad.1569307.
[5] J. Bassler et al., J. Cell Biol. 207, 481–98 (2014); doi: 10.1083/jcb.201408111.
[6] O. Svensson, et al., Acta Crystallogr. D. Biol. Crystallogr. 71, 1757–67 (2015); doi: 10.1107/S1399004715011918.
[7] M.W. Bowler, et al., J. Synchrotron Radiat. 22, 1540–7 (2015); doi: 10.1107/S1600577515016604.
Top image: Structure of the Aspergillus nidulans Rpf2-Rrs1 complex.