- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2011
- Structural biology
- The Mre11/Rad50 complex - an ATP dependent molecular clamp in DNA double strand break repair
The Mre11/Rad50 complex - an ATP dependent molecular clamp in DNA double strand break repair
The DNA in our cells is continuously exposed to DNA damaging agents such as radiation and mutagenic chemicals. Amongst the various forms of damage, probably the most cytotoxic are DNA double strand breaks (DSBs) which can lead to gross chromosomal aberrations and cancer. DSBs arise during chromosomal replication but can also be products of ionising radiation and genotoxic chemicals. In addition, enzymatically induced DSBs are intermediates in V(D)J recombination, immune globulin class switching and meiosis. Both sporadic and programmed DSBs need careful and prompt repair to maintain genome stability and allow cell survival.
DSBs elicit a complex cellular response and induce predominantly two different repair pathways: homologous recombination (HR) and non-homologous end joining (NHEJ). In HR, DNA ends are resected and paired with the sister chromatid to prime repair synthesis, resulting in error-free repair. In contrast, in NHEJ the DNA ends are directly ligated without a template, making NHEJ potentially mutagenic.
The Mre11/Rad50/Nbs1 complex (MRN; Mre11/Rad50/Xrs2 in Saccharomyces cerevisiae) is an evolutionarily conserved key player in the detection and repair of DNA double strand breaks. Mutations in MRN subunits lead to a defective DSB repair response and can result in cell death or chromosomal aberrations. Hypomorphic mutations in the MRN-ATM axis are linked to diseases that are characterised by neurological disorders and/or cancer predisposition. The MRN complex functions in both the HR and NHEJ repair pathways and is involved in DSB detection, DNA tethering, damage signalling via ATM, structural organisation of repair and checkpoint foci, as well as nucleolytic processing of the DNA ends. These diverse functions suggest that the MRN complex possesses multiple activities, including not only enzymatic, but also architectural and structural roles. Whereas enzymatic activities are required to prepare DNA ends for repair, architectural functions maintain these in close proximity. Homologs of the Mre11/Rad50 complex are found in every kingdom of life, suggesting that this core enzyme complex is fundamental for genomic stability. To understand the interplay between Mre11 and Rad50, we determined the crystal structure of the catalytic head of the Thermotoga maritima (Tm) Mre11/Rad50 complex (SbcCD).
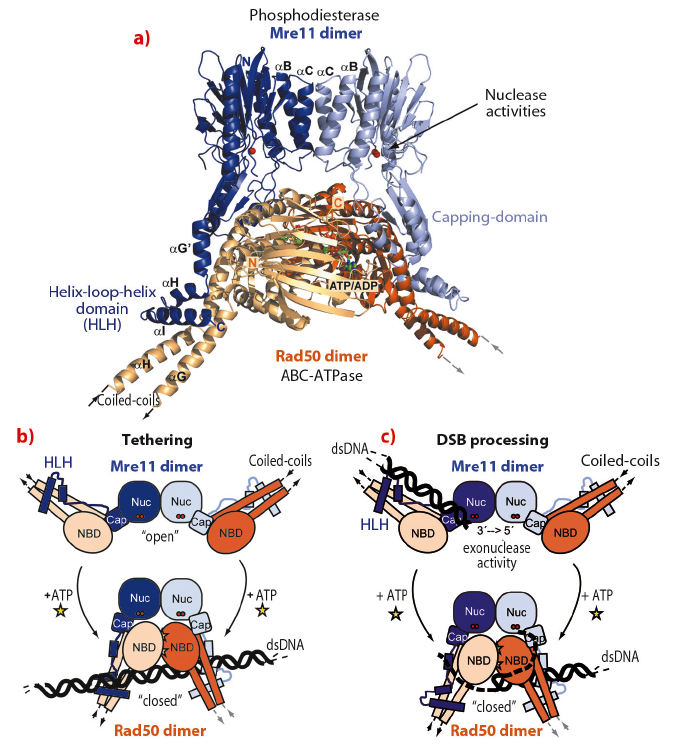
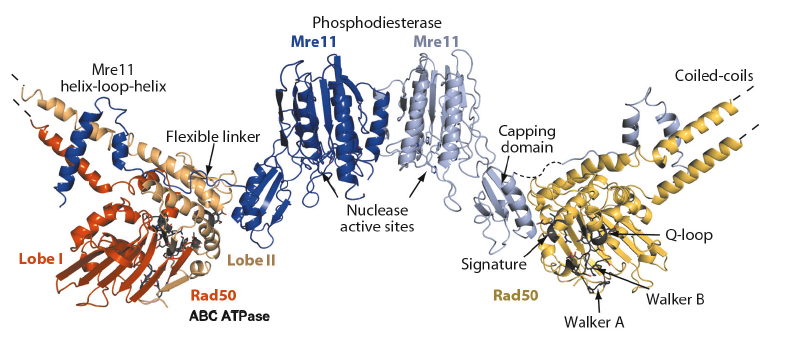
The structure reveals important information about the architectural organisation of Mre11 homodimerisation and the interaction of Mre11 with Rad50. The TmMR catalytic head (TmMRNBD) is an elongated crescent shaped complex and its core is formed by a dimer of the two Mre11 nuclease domains, with the two nuclease active sites located near the centre of the concave face (Figure 99). The Rad50 NBDs each attach to the outside of the nuclease dimer and form the tips of the crescent. The Rad50 coiled-coils protrude from the convex side of the catalytic head, opposite to the nuclease active sites, at an angle of approx. 120° from each other. The overall structure explains two poorly understood functional characteristics of MR. The widely separated, outward placement of the Rad50 NBDs allows unobstructed access of DNA to the Mre11 active sites, even if the DNA ends are blocked by large proteins. In addition, our structure together with SAXS and chemical crosslinking analysis of the MR head complex, with and without nonhydrolysable ATP analogues, shows that the complex undergoes major conformational changes upon ATP binding to Rad50 [1].
 |
|
Fig. 99: Overall structure of the Mre11:Rad50 catalytic head complex in its “open” nucleotide free conformation. |
Recently we determined the crystal structure of the closed ATP bound TmMre11/Rad50 head complex (TmMRNBD ATP) and on the basis of both structures we have derived a precise structural framework for ATP induced conformational changes in the TmMRNBD complex. We find that ATP binding induces a large structural change from the open form with accessible TmMre11 nuclease sites into a closed form (Figure 100a). Most remarkably, ATP repositions the engaged Rad50 NBDs into the dsDNA binding groove of the Mre11 active site, suggesting an ATP induced blocking of Mre11’s active site. While this conformation is incompatible with dsDNA binding to Mre11, unwound or single-stranded DNA may still reach Mre11’s nuclease sites and could offer a structural explanation of how MR promotes endonucleolytic cleavage of single-stranded DNA or hairpins (Figure 100b). Additionally, the extended coiled-coil domains of Rad50 undergo an extensive conformational reorientation, which could prevent intramolecular MR interaction after ATP dependent engagement and allow intermolecular interaction between different MR complexes via the zinc hook. This raises the possibility that DNA tethering functions of the complex and ATP dependent DNA binding are allosterically coupled (Figure 100c).
Principal publication and authors
K. Lammens (a,b), D. Bemeleit (b), C. Möckel (b), E. Clausing (b) A. Schele (b), S. Hartung (b), C.B. Schiller (b), M. Lucas (b), C. Angermüller (b), J. Söding (b), K. Sträßer (b) and K.P. Hopfner (a,b), Cell 145, 54-66 (2011); C. Möckel (b), K. Lammens (a,b), A. Schele (b) and K.P. Hopfner (a,b), Nucleic Acids Res. 39, 1-14 (2011).
(a) Center for Integrated Protein Science Munich (CIPSM)
(b) Gene Center, Ludwig-Maximilians-University Munich (Germany)
References
[1] C. Wyman, J. Lebbink and R. Kanaar, DNA Repair 10, 1066-1070 (2011).