- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2011
- Structural biology
- Structures of SAS-6 suggest its organisation in centrioles
Structures of SAS-6 suggest its organisation in centrioles
Centrioles are nine-fold symmetric, cylinder-shaped organelles that organise centrosomes and that are strictly required as a template for cilia and flagella. Abnormalities in these structures can result in numerous human diseases like ciliopathies or infertility. Thus, a detailed understanding of centriole architecture and formation is an important goal [1].
The diameter of centrioles and their symmetry is determined early on in their assembly by a so-called cartwheel structure [2]. This cartwheel consists of a central ring-like hub and nine spokes that radiate outwards to the periphery of centrioles where centriolar microtubule triplets (Figure 117a) are found. Through an elongation process and further additions of components, mature centrioles are subsequently formed. Since the cartwheel structure provides a scaffold involved in the establishment of the symmetry and diameter of centrioles, understanding the structural organisation of cartwheels should reveal important aspects of centriole formation. A prime candidate for a key player in this respect is the centriolar protein SAS-6. SAS-6 is found at the hub of the cartwheel and is required for its formation. It is found in all organisms that have centrioles and, in all systems tested so far, is essential for proper centriole formation [2].
 |
|
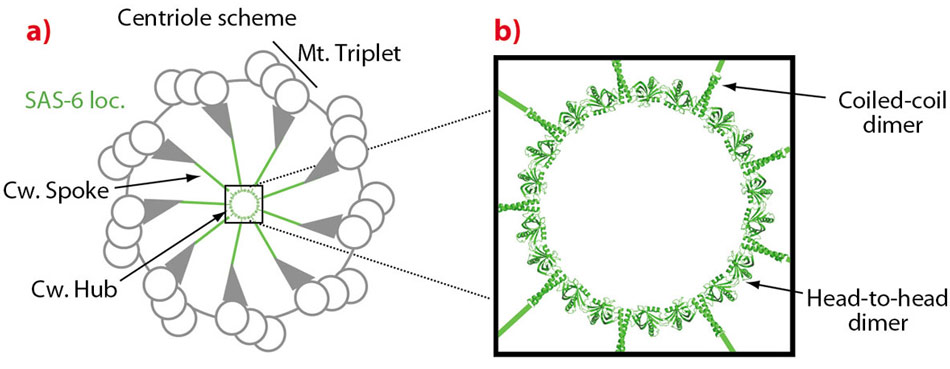
Fig. 117: a) Centriole scheme with peripheral microtubule triplets (Mt. Triplet) and central cartwheel structure (Cw.) with hub and spokes. In green is the location of SAS-6. A 9-fold symmetric SAS-6 ring model is placed at the cartwheel hub. b) A close-up view of the cartwheel hub. Solid green lines indicate that the SAS-6 coiled-coil domain could, theoretically, protrude to the cartwheel periphery if it is fully extended. |
To understand how SAS-6 might organise the centriolar cartwheel, we obtained high-resolution crystal structures of SAS-6 fragments at beamlines ID29 and BM14. These structures show that SAS-6 consists of a globular N-terminal head domain and a rod like coiled-coil domain that follows in sequence. Both domains were found to form homodimers: The N-terminal domain formed a slightly curved head-to-head dimer while the coiled-coil domain formed a canonical elongated coiled-coil dimer. We confirmed these interactions in solution and showed that the coiled-coil dimer is stable while head-to-head dimerisation occurs with a rather low affinity. Both interactions are biologically relevant and are essential for SAS-6 function in vivo.
Modelling these interactions in SAS-6 resulted in curved oligomers that were compatible with a 9-fold ring with similar dimensions to that of cartwheel hubs observed in vivo (Figure 117b). Consistent with this model, we found that recombinant SAS-6 constructs do indeed form higher-order oligomers in solution and that they, as judged by cryo-EM, can adopt structures similar to centriolar cartwheels.
In summary, we have shown that, through self-association, a single protein – SAS-6 – can account for the organisation of centriolar cartwheel centres and thereby contribute to the establishment of centriolar symmetry and dimensions. The challenge ahead is to determine how the other key components of centriole assembly contribute to this process. High-resolution crystal structures of these components will undoubtedly play a decisive role in this.
Principal publication and authors
M. van Breugel (a), M. Hirono (b), A. Andreeva (a), H.-a. Yanagisawa (b), S. Yamaguchi (b), Y. Nakazawa (b,d), N. Morgner (c), M. Petrovich (a), I.-O. Ebong (c), C.V. Robinson (c), C.M. Johnson (a), D. Veprintsev (a,e) and B. Zuber (a), Science 331, 1196 (2011).
(a) Medical Research Council – Laboratory of Molecular Biology, Cambridge (UK)
(b) Department of Biological Sciences, University of Tokyo (Japan)
(c) Chemistry Research Laboratory, University of Oxford (UK)
(d) Current Address: Department of Biosciences, School of Science, Kitasato University, Sagamihara (Japan)
(e) Current Address: Paul Scherrer Institut, Villigen (Switzerland)
References
[1] E.A. Nigg and J.W. Raff, Cell 139, 663 (2009).
[2] P. Strnad and P. Gonczy, Trends Cell Biol 18, 389 (2008).



