- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2000
- Life Sciences
- The Structure of the 30S Ribosome Subunit
The Structure of the 30S Ribosome Subunit
In all organisms, the information present in the gene is copied to an intermediate molecule called messenger RNA (mRNA), and then translated into protein by the ribosome. The process is mediated by an adapter molecule called tRNA, which contains an amino acid linked at one end, with a three-base anti-codon at the other end, which is complementary to the codon on the gene. Ribosomes are large nucleoprotein particles, and consist of two subunits, denoted small (or 30S) and large (or 50S) in bacteria. They have a mass of about 2.5 MDa, and are about 2/3 RNA by mass. Eukaryotes and archaea have significantly larger ribosomes but are thought to function in essentially the same way. The 50S subunit contains the site of peptidyl-transferase activity, which catalyses the formation of the peptide bond during the addition of an amino acid to the end of a growing polypeptide chain. The 30S subunit provides the binding site for mRNA and is responsible for monitoring base-pairing between the codon on mRNA and the anticodon on tRNA. Ribosomes are also the target for a large number of clinically important antibiotics, including streptomycin, gentamycin, erythromycin and tetracycline. However, despite four decades of biochemical work, details of ribosome function and the action of antibiotics have been unclear due to lack of structural information. The ribosome is a very large RNA-protein complex, so its structure is also extremely important for understanding RNA structure and its interactions with proteins.
Until recently, structural studies on ribosomes were hindered by difficulties in obtaining crystals that diffracted to high resolution. With respect to the 30S subunit, it is now possible to obtain, in a regular manner, crystals that diffract to 3.0 Å [1]. However, key technological developments were also required and these include (a) highly monochromatic, tuneable and brilliant synchrotron beamlines made possible by insertion devices, (b) fast, large and high-resolution area detectors, (c) fast computing, and (d) better crystallographic software. Another essential advance was the development of cryocrystallography.
Several approaches have been used to obtain phase information, including the use of heavy atom clusters, cryoelectron microscopic reconstructions as starting models for low-resolution molecular replacement, or the use of anomalous scattering. In our case, an initial 5.5 Å structure was obtained by phases from a combination of heavy atom clusters and anomalous scattering [1]. In hindsight, it appears that the heavy atom cluster phases were not essential, though they were extremely useful in the identification of heavy atom sites.
To obtain high resolution phases (to about 3.2 Å), we relied heavily on anomalous scattering from LIII edges. These experiments were mainly done at the APS SBC ID-19 beamline and also at the ID14-4 beamline at the ESRF. The LIII edges of lanthanides and osmium have a very strong white line, which gives rise to about 20-30 electrons of anomalous scattering per atom. By orienting crystals and measuring Bijvoet pairs accurately, we were able to get maps that were sufficient to build an atomic structure and refine it to a resolution of 3 Å [2]. We have also solved the structure of the 30S in complex with three antibiotics, streptomycin, paromomycin and spectinomycin, using data collected at ID14-4 [3].
The structure is shown in Figure 6. It consists of a 1522 nucleotide piece of RNA and 20 polypeptide chains. Using the structure, we have been able to analyse the binding sites for tRNA and mRNA. We were helped by the fact that a stem-loop of RNA from a neighbouring molecule in the crystal is inserted into one of the tRNA binding sites. The end of ribosomal RNA is also folded back into the mRNA binding cleft, so the structure gives a clear idea of the path of mRNA in the 30S. The antibiotics themselves shed light not only on how they work, but on several crucial ribosome functions such as translocation, and decoding, or how the 30S discriminates between correct and incorrect tRNAs based on whether they are complementary to the codon on mRNA or not.
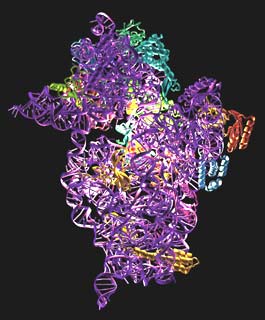 |
Fig. 6: The structure of the 30S subunit. The RNA is shown in purple and the proteins in various colours. The binding site for the antibiotics is highlighted.
|
These structures, along with other ribosome structures from other groups, will greatly accelerate the process of understanding translation at a molecular level.
References
[1] W.M. Clemons, J.L.C. May, B.T. Wimberly, J.P. McCutcheon, M. Capel and V. Ramakrishnan, Nature 400, 833-840 (1999).
[2] B.T. Wimberly, D.E. Brodersen, W.M.J. Clemons, R. Morgan-Warren, C. von Rhein, T. Hartsch and V. Ramakrishnan, Nature 407, 327-339 (2000).
[3] A.P. Carter, W.M.J. Clemons, D.E. Brodersen, R. Morgan-Warren, B.T. Wimberly, and V. Ramakrishnan, Nature 407, 340-348 (2000).
Authors
D.E. Brodersen, A.P. Carter, W.M. Clemons Jr., B.T. Wimberly, and V. Ramakrishnan.
MRC, Cambridge (UK)



