- Home
- News
- Spotlight on Science
- Liquid-liquid phase...
Liquid-liquid phase separation of Rubisco during carboxysome biogenesis in cyanobacteria
14-05-2019
Cyanobacteria sequester the enzyme Rubisco into carboxysomes for efficient carbon fixation during photosynthesis. Assembly of these organelles is not well understood. Crystal structures obtained at the ESRF helped in understanding how the adapter protein CcmM concentrates Rubisco into a dynamic, liquid-like network.
Essentially all organic material on earth is the product of photosynthesis, the process that channels the energy from sunlight into the fixation of carbon from atmospheric carbon dioxide. Cyanobacteria are reported to be the first organisms on Earth to fix CO2 via oxygen producing photosynthesis and today account for ~10% of worldwide CO2 fixation. The key enzyme performing carbon fixation is Rubisco (Ribulose-1,5-bisphosphate carboxylase/oxygenase), a ~520 kDa complex of eight large (RbcL) and eight small (RbcS) subunits. Rubisco catalyses the carboxylation of the 5-carbon sugar substrate ribulose-1,5-bisphosphate. However, the enzyme is rather inefficient, with a slow turnover rate and limited selectivity for CO2 versus O2. To overcome this problem, cyanobacteria have evolved a CO2 concentrating mechanism (CCM) by sequestering Rubisco together with the enzyme carbonic anhydrase (CA) into microcompartments called carboxysomes. This serves to generate high levels of CO2 in the vicinity of Rubisco, thus allowing cyanobacteria to use less selective Rubiscos that are ~10 times faster than the enzymes of land plants. Substantial gains in productivity could be achieved by transplanting functioning carboxysomes into the chloroplasts of crop plants.
Carboxysomes resemble intracellular viruses with a highly symmetric proteinaceous shell, and have been classified into α and β based on the type of Rubisco they encapsulate. During β-carboxysome biogenesis, Rubisco first aggregates, mediated by the protein CcmM, followed by shell formation. There are two isoforms of CcmM in the cyanobacterium Synechococcus elongatus PCC 7942. The full-length protein (also called M58) contains a CA-like domain, followed by three small subunit-like (SSUL) modules that have 25% sequence identity to RbcS of Rubisco. A smaller isoform, called M35, consists only of the three SSUL modules. It had been speculated that these domains might replace RbcS subunits, thereby linking Rubisco complexes into a 3D network [1-3].
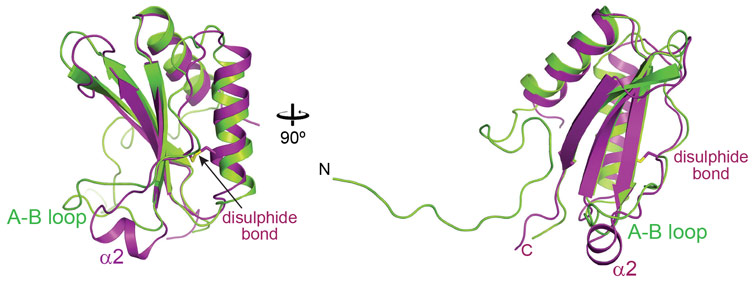
Crystal structures of the SSUL module were solved with X-ray diffraction data collected at beamlines ID23-2 and MASSIF1, and showed that the SSUL backbone indeed resembled Rubisco small subunits (Figure 1). A surprising feature was the presence of an intramolecular disulphide bond between conserved cysteine residues, suggesting possible redox regulation. Moreover, the SSUL structure showed a short helical insertion, α2, in place of the A-B loop in RbcS (Figure 1). Biochemical binding assays demonstrated that Rubisco aggregation by M35 required the fully-assembled Rubisco and did not occur with assembly intermediates lacking RbcS. Reduced M35 was ~5-fold more efficient in binding than the oxidised protein. This is consistent with Rubisco aggregation occurring in the reducing environment of the cytosol, whereas the interior of the carboxysome is oxidixing. Further analysis by fluorescence recovery after photobleaching (FRAP) showed that M35 induced the phase separation of Rubisco into a liquid-like network, which was more dynamic under oxidising conditions. Prevention of disulphide-bond formation in vivo resulted in abnormal carboxysomes, a ~20-fold increase in CO2 requirement and a ~4-fold slower growth rate of mutant cyanobacteria, supporting the view that redox-regulation in the SSUL module is critical for carboxysome biogenesis and function.
 |
|
Figure 1. Overlay of the crystal structures of the oxidised SSUL1 module of CcmM from Synechococcus elongatus PCC7942 (in magenta) and the small subunit of Rubisco, RbcS (in green). |
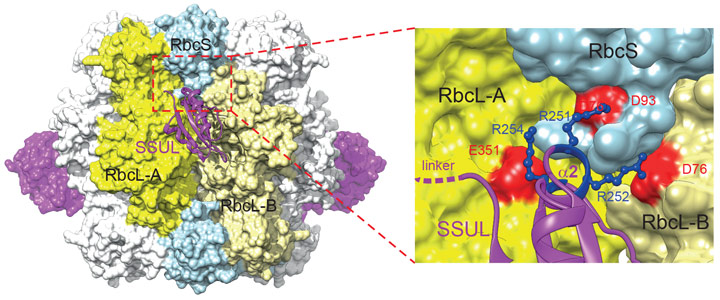
The molecular organisation of the M35-Rubisco network within liquid droplets was analysed by cryo-electron microscopy. The structure of the M35-Rubisco complex at 2.8 Å resolution revealed four pairs of symmetry-equivalent SSUL binding sites at the equator of Rubisco (Figure 2). Interestingly, the symmetry-related binding sites overlap so that only a maximum of four SSUL modules can dock. Contrary to the earlier model, the SSUL modules do not displace RbcS but in fact require the presence of RbcS for binding, with critical contacts mediated by the helix α2 of SSUL (Figure 2). This mechanism ensures that only Rubisco holoenzyme is packaged into carboxysomes.
Taken together, the findings suggest a model in which Rubisco and CcmM form a liquid-like, phase-separated condensate into which other carboxysome components can exchange. Upon shell formation, preventing entry of reducing agents, the interior of the carboxysome becomes oxidising and renders the Rubisco-CcmM interaction more dynamic. The data show why the CcmM protein is an indispensable component of the carboxysome assembly machinery, providing valuable insights for ongoing attempts to introduce carboxysomes into plant chloroplasts. More efficient carbon fixation would help to meet the food security demands for an ever-growing human population.
Principal publication and authors
Rubisco condensate formation by CcmM in β-carboxysome biogenesis, H. Wang (a), X. Yan (a), H. Aigner (a), A. Bracher (a), N.D. Nguyen (b), W.Y. Hee (b), B.M. Long (b), G.D. Price (b), F.U. Hartl (a) & M. Hayer-Hartl (a), Nature 566, 131 (2019); doi: 10.1038/s41586-019-0880-5.
(a) Max Planck Institute of Biochemistry, Martinsried (Germany)
(b) The Australian National University, Canberra (Australia)
References
[1] G.S. Espie, and M.S. Kimber, Photosynth. Res. 109, 7-20 (2011).
[2] B.D. Rae, B.M. Long, M.R. Badger, and G.D. Price, Microbiol. Mol. Biol. Rev. 77, 357-379 (2013).
[3] C.A. Kerfeld, and M.R. Melnicki, Curr. Opin. Plant Biol. 31, 66-75 (2016).
Top image: Model of the Rubisco-SSUL complex showing a hypothetical alternating arrangement of four SSUL modules (magenta) bound.