- Home
- News
- Spotlight on Science
- The remarkable complexity...
The remarkable complexity of vitamin B6 biosynthesis
13-04-2017
Vitamin B6 biosynthesis has been studied with X-ray crystallography and in crystallo spectroscopy. An unprecedented use of imine chemistry creates covalently-linked intermediates that allow migration of the reaction across a path of over 20 Å between substrate- and product-binding sites.
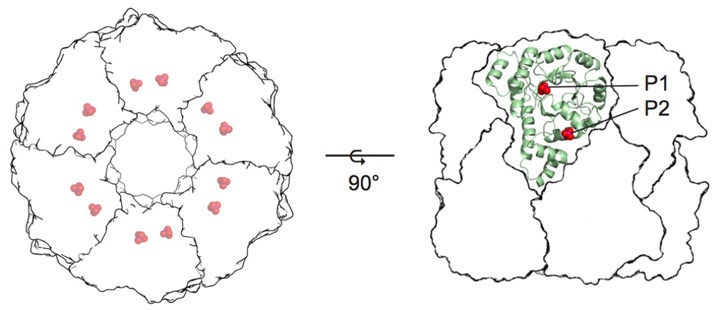
The mechanism of vitamin B6 biosynthesis has long been enigmatic [1]. The main biosynthetic route uses an enzyme complex known as pyridoxal 5-phosphate (PLP) synthase. The full PLP synthase complex consists of twelve copies each of the Pdx1 and Pdx2 proteins (Figure 1) [2]. While Pdx2 simply catalyses hydrolysis of glutamine to generate the ammonia required for PLP biosynthesis, Pdx1 catalyses the condensation of two carbohydrate substrates and ammonia in a complex reaction involving more than ten catalytic steps.
 |
|
Figure 1. Crystal structure of pyridoxal 5-phosphate synthase. P1 and P2 indicate phosphate-binding sites. |
The Pdx1 enzyme has a (βα)8-barrel fold and forms a dodecamer. Each Pdx1 enzyme contains two phosphate-binding sites, named P1 and P2 that are separated by 21 Å (phosphorus to phosphorus) [1]. The first Pdx1 substrate is ribose 5-phosphate (R5P), which binds with its phosphate group in the P1 site [3,4]. Addition of the second Pdx1 substrate, ammonia, leads to the formation of the chromophoric I320 intermediate with an absorbance maximum λ = 320 nm [5]. The formation of I320 partially vacates the P1 active site and creates space for the third substrate, glyceraldehyde 3-phosphate (G3P), making the P1 site a dual specificity active site. As the product, PLP, is observed with its phosphate group bound in the P2 site, the bridging position of the I320 intermediate between P1 and P2 sites explains intermediate transfer, a novel example of channelling.
The structures reported are enzyme-intermediate complexes that form the basis for mechanistic proposals of PLP biosynthesis. The data were collected over nearly a decade, testing several Pdx1 enzymes from different organisms. More than 1,000 crystals were tested to optimise soaking protocols used to provide both high-resolution diffraction and homogeneous accumulation of Pdx1 intermediates.
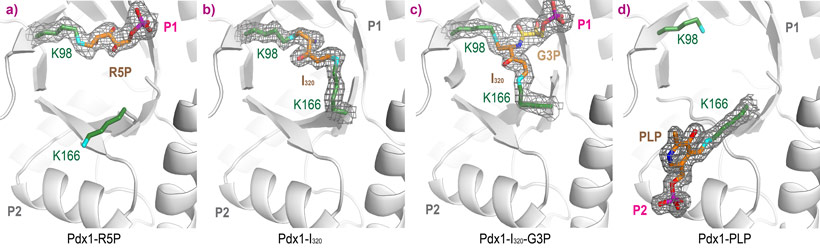
Binding of R5P occurs by covalent attachment through Schiff base formation with Lys98 (Figure 2a, collected at ESRF ID14-1, 1.9 Å resolution). Addition of ammonia to Pdx1-R5P complexes leads to formation of I320 through a second Schiff base with Lys166 (Figure 2b, collected at Diamond I04-1, 1.7 Å resolution). Adding G3P to Pdx1-I320 leads to a covalent complex with the G3P phosphate bound in the P1 site (Figure 2c, collected at ESRF ID23-1, 1.9 Å resolution). In the product complex, PLP is covalently bound through Schiff base formation with Lys166, with its phosphate bound in the P2 site (Figure 2d, collected at ESRF ID14-1, 1.6 Å resolution).
 |
|
Figure 2. Complexes of Pdx1 after sequential addition of a) ribose 5-phosphate R5P, b) ammonia and c) glyceraldehyde 3-phosphate G3P, leading to the formation of d) pyridoxal 5-phosphate. |
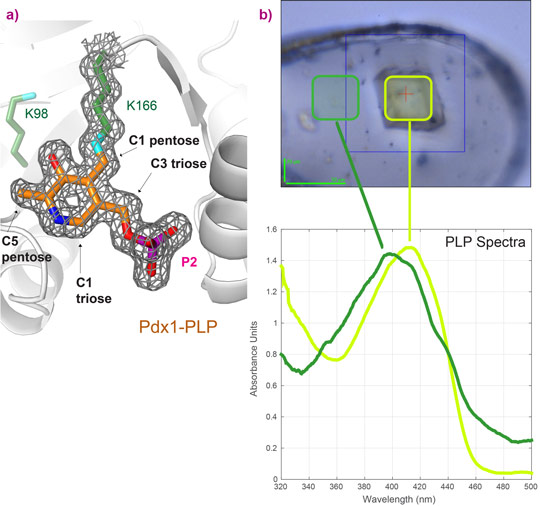
The use of micro-spectrophotometry at the ESRF Cryobench was essential to show that crystals used for data collection contained the desired chromophoric intermediates. We have used online spectroscopy to provide proof of the chemical nature of certain intermediates, such as PLP (Figure 3); the spectrum of a crystal with PLP bound shows an absorption maximum at λ = 414 nm characteristic for covalently bound PLP, light green, while free PLP in the surrounding buffer has an absorption maximum of λ = 388 nm, dark green (Figure 3b, collected at ESRF ID14-1 using on-line micro-spectrophotometry).
 |
|
Figure 3. Confirmation of the covalent PLP-enzyme complex by a) structure determination and b) optical spectroscopy at the Cryobench. |
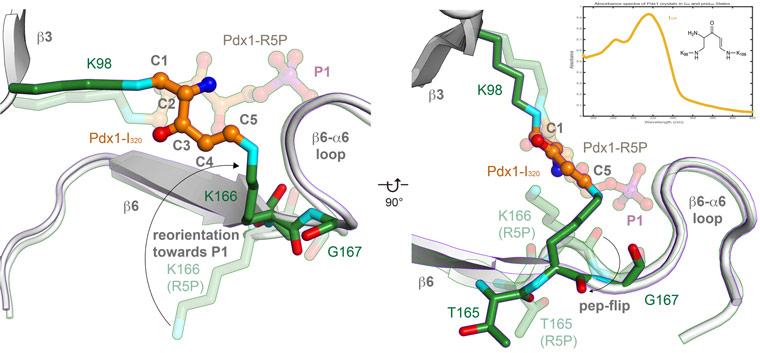
Spectrophotometry can also be used to monitor specific radiation damage of intermediates, as was done to monitor specific radiation damage of the I320 intermediate (Figure 4, collected on-line at ESRF ID14-4). Determination of a low-dose structure of the I320 complex used about 50 complete data sets for I320 complexes (collected at ESRF ID23-1). Data collected before the 245 kGy threshold (corresponding to 80% retained absorbance of the I320 species, not shown) were merged after identifying groups that merged well with the software BLEND, and the final dataset contained 19-wedges from nine crystals to confirm the structure of the I320 intermediate as a covalently linked intermediate between two lysine residues.
 |
|
Figure 4. The central intermediate of PLP biosynthesis is the doubly covalently linked I320. |
Pdx1 uses covalent tethers in the synthesis of PLP to prevent loss of intermediates to surrounding solvent. This strategy maintains a high local concentration of substrate and protects the reactive I320 species. Parallels exist to the phosphopantothenoyl thioesters that are used to transfer intermediates between active sites in the assembly line enzymology of fatty acid synthase, polyketide synthases and non-ribosomal polypeptide synthases. Pdx1 uses a similar transfer strategy to those found in the glycine cleavage system and the classical pyruvate dehydrogenase complex, where lipoic acid is used to chaperone intermediates between active sites. While the use of covalent tethers is common in all these examples, Pdx1 transfers covalent intermediates within a single catalytic domain. The intricate relay mechanism displayed by the Pdx1 subunit of PLP synthase allows the enzyme to maintain precise control of the complex reaction performed across multiple active sites.
Principal publication and authors
Lysine relay mechanism coordinates intermediate transfer in vitamin B6 biosynthesis, M.J. Rodrigues (a,b), V. Windeisen (a,c), Y. Zhang (d), G. Guédez (c), S. Weber (c), M. Strohmeier (c), J.W. Hanes (d,e), A. Royant (f,g), G. Evans (b), I. Sinning (c), S.E. Ealick (d), T.P. Begley (h) and I. Tews (a), Nat Chem Biol. 13, 290-294 (2017); doi: 10.1038/nchembio.2273.
(a) Biological Sciences, University of Southampton, Southampton (UK)
(b) Diamond Light Source, Harwell Science and Innovation Campus, Didcot (UK)
(c) Heidelberg University Biochemistry Center (BZH), Heidelberg (Germany)
(d) Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York (USA)
(e) Pacific Biosciences, Menlo Park, California (USA)
(f) Institut de Biologie Structurale, Université Grenoble Alpes, CNRS, CEA, Grenoble (France)
(g) ESRF
(h) Department of Chemistry, Texas A&M University, College Station, Texas (USA)
References
[1] T.B. Fitzpatrick et al., Biochem J. 407, 1-13 (2007).
[2] M. Strohmeier et al., PNAS 103, 19284-9 (2006).
[3] A.M. Smith et al., J Biol Chem. 290, 5226-39 (2015).
[4] G. Guédez et al., Structure 20, 172-84 (2012).
[5] G.C. Robinson et al., PNAS 113, E5821-9 (2016).



