- Home
- News
- Spotlight on Science
- Sub-angstrom resolution...
Sub-angstrom resolution protein crystallography reveals how living organisms discriminate in favour of phosphate in arsenate-rich environments
05-03-2013
Arsenate and phosphate are both common on Earth and have very similar chemical properties. However, phosphate is vital to life while arsenate is toxic. Discrimination by proteins between arsenate and phosphate is thus essential to the survival of living organisms, particularly those found in environments rich in arsenate. A comparison of the ultra high resolution crystal structures of a phosphate binding protein in complex with either phosphate or arsenate has allowed the elucidation of the mechanism for phosphate discrimination and revealed this to be based on the geometry of negative-charge-assisted hydrogen bonds formed between arsenate and phosphate anions and the protein.
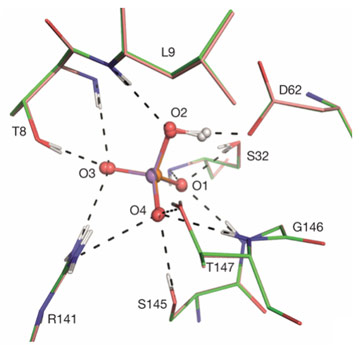
The mechanism of discrimination by proteins between phosphate and arsenate - one vital to the survival of living organisms, the other highly toxic - has thus far been unclear due to the very high level of similarity of the chemical properties of the two anions, including their pKa [1], number and charge of oxygen atoms and thermochemical radii [2]. The discrimination mechanism has now been elucidated as a result of a comparison of the ultra-high resolution crystal structures of the phosphate binding protein (PBP) from Pseudomonas fluorescens SBW25 in complex with either phosphate (PO42-) or arsenate (AsO42-) at ph 4.5 and 8.5. The crystal structures (Figure 1) were obtained using diffraction data collected at ESRF beamlines ID29 and ID23-1. PBPs are components of the phosphate-specific transport systems found in bacteria and archea and which are activated at low phosphate or high arsenate conditions [3]. PBPs bind phosphate with sub-micromolar Kds and discriminate between phosphate and arsenate by factors of between 500 and 850.
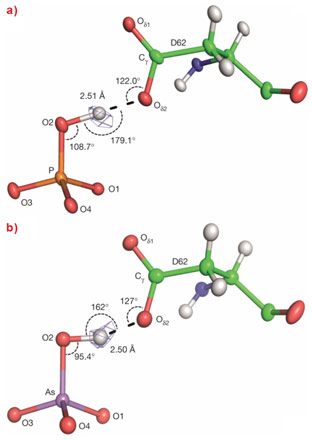
The sub-angstrom resolution obtained from the diffraction data allowed the location - albeit at low signal-to-noise levels - of hydrogen atoms in the electron density maps. These confirm, as expected from pKa values, that at both pH values both anions are bound in their dibasic forms (i.e. H(As/P)O42-) and, at first sight, the binding modes of the two moieties to the protein appear identical (Figure 1). Nevertheless, closer inspection reveals a difference in the geometries of the short (d ~2.5 Å) low-barrier hydrogen bonds - more specifically, heteromolecular negative-charge-assisted hydrogen bonds ((-)CAHBs) - linking the side chain carboxyl group of Asp 62 to O2 of the phosphate or arsenate anions (Figure 2). In the PBP-HPO42- complex, the geometry of this low-barrier hydrogen bond is close to the canonical values expected for an optimal interaction (P–O2–H, 108.7°; O2–H•••Oδ25, 179.1°; Cγ-Oδ2,•••H 122°). However, although the thermochemical radii of HPO42- and HAsO42- differ by only 4% [2], the longer As-O2 bond length leads to a distortion of the topology of the equivalent (-)CAHB (As–O2–H, 95.4°; O2–H•••Oδ25, 162°; Cγ-Oδ2,•••H 127°) in the PBP-HAsO42- complex.
It thus appears that a small, very subtle difference in hydrogen bonding geometry is the main determinant of the extremely high degree of phosphate/arsenate selectivity of the PBPs of organisms found in arsenate-rich environments. This subtle difference in hydrogen bonding geometry was visible only thanks to ultra-high resolution diffraction data that is increasingly available at the ESRF's high intensity macromolecular crystallography beamlines.
* Omit Fourier difference maps are created by including a ligand/atom in the model, refining the structure, removing what was added, re-refining the structure, then calculating a difference Fourier map. This should only show the added ligand/atom, if the model is correct.
Principal publication and authors
The molecular basis of phosphate discrimination in arsenate-rich environments, M. Elias (a), A. Wellner (a), K. Goldin-Azulay (a), E. Chabriere (b), J.A. Vorholt (c), T.J. Erb (c), D.S. Tawfik (a), Nature 491, 134-137 (2012).
(a) Department of Biological Chemistry, Weizmann Institute of Science, Rehovot (Israel)
(b) Faculté de Médecine et de Pharmacie, CNRS-Université de la Méditerranée, Marseille (France)
(c) Institute of Microbiology, Eidgenössische Technische Hochschule, Zurich (Switzerland)
References
[1] R.A. Kohn & T.F. Dunlap, J. Anim. Sci. 76, 1702–1709 (1998).
[2] M.M. Kish, & R.E. Viola, Inorg. Chem. 38, 818–820 (1999).
[3] G.R. Willsky, & M.H. Malamy, J. Bacteriol. 144, 366–374 (1980).
Top image: Discrimination is provided by the geometry of a hydrogen bond formed between an oxygen of phosphate or arsenate and a carboxyl group of the protein.