- Home
- News
- Spotlight on Science
- A Molecular Machine...
A Molecular Machine for Processing DNA Breaks
28-01-2005
All living organisms are faced with the challenge of maintaining the integrity of their genetic information, the DNA genome. This is under constant assault from a wide variety of internal and external factors and cells have evolved sophisticated ways to repair damage such as double-strand breaks in the DNA [1].
One such method of DNA repair is the process of homologous recombination (HR). This involves copying information from an identical (homologous) undamaged section of DNA, and 'splicing' the corrected sequence into the damaged strand of DNA. The crystal structure of a key player in this process, the RecBCD multi-enzyme complex bound to DNA has recently been solved by M.R. Singleton et al. [2] using X-rays from beamlines ID14-1 and ID14-4.
The RecBCD complex rapidly binds the ends of DNA resulting from double strand breaks, and carries out the so-called resectioning process. This involves an unwinding of the DNA and digestion of DNA strands to leave what is known as a Chi-terminated tail. The enzyme then loads the strand exchange factor RecA onto the tail in preparation for the next stage of the process.
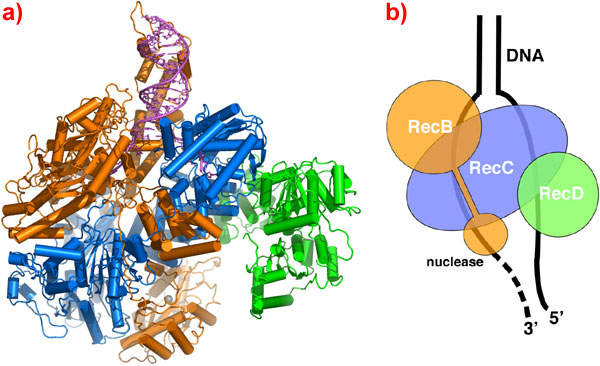
Fig. 1: (a) Ribbon representation of the RecBCD complex. RecB is shown in orange, RecC in blue and RecD in green. The DNA is shown in magenta. The nuclease active site has a calcium ion bound in it, shown as a grey sphere. (b) Schematic diagram of the complex showing the path taken by the DNA through the complex.
The crystal structure (Figure 1a) shows the architecture of the complex. RecC forms the core of the complex with RecB wrapped around it and RecD sitting below it. RecB and RecD are helicase proteins, 'molecular motors' which cooperate to unwind the DNA and feed the separated strands through channels in RecC. The 3' strand emerges from RecC straight into the nuclease domain of RecB (Figure 1b) where it is digested. RecC also contains a DNA-scanning function which looks for the Chi sequence as DNA spools through. When it does an, as-yet undetermined, conformational change switches the nuclease to the other strand and initiates RecA loading.
M.R. Singleton kindly acknowledges funding by 'Cancer Research, UK'
References
[1] S.C. Kowalczykowski, Initiation of genetic recombination and recombination-dependent replication. Trends. Biochem. Sci. 25, 156-165 (2000).
[2] M.R. Singleton, M.S. Dillingham, M. Gaudier, S.C. Kowalczykowski, and D.B. Wigley, Crystal structure of RecBCD enzyme reveals a machine for processing DNA breaks, Nature 432, 187-193 (2004).
Authors
M.R. Singleton (a), M.S. Dillingham (b), M. Gaudier (a), S.C. Kowalczykowski (c) and D.B. Wigley (a).
(a) Cancer Research UK, Clare Hall Laboratories, South Mimms (U.K.)
(b) National Institute for Medical Research, Mill Hill (U.K.)
(c) Sections of Microbiology and Molecular & Cellular Biology, Center for Genetics and Development, University of California at Davis (U.S.A.)