- Home
- News
- Spotlight on Science
- Cell guidance mechanisms...
Cell guidance mechanisms revealed at atomic-level resolution
28-10-2010
The mechanism underlying the interaction between two proteins that act to position cells, for example in the growth and guidance of developing nerve fibres in the brain, has been revealed by a new collaborative study between researchers at the University of Oxford and Trinity College Dublin.
The assembly of the circuitry of the brain is one of the most impressive feats in the development of an organism. The brain is composed of thousands of distinct regions, which must be wired together in a very precise fashion to enable them to carry out their specific functions. Each area may contain millions of individual cells, each of which has to find its way to the correct position in the brain and project its nerve fibre (called an axon) to its appropriate targets. This is accomplished as growing axons sense and respond to cues on other cells in the tissue of the developing brain, which control the direction of projection of each axon.
The outer membrane of each cell is studded with protein molecules which act both as sensors, or receptors, for these cues and as cues for other cells to respond to. One important class of proteins which has been known for some time to be involved in guiding growing axons is the family of “semaphorins” – named in analogy of the signalling system between ships at sea. One of these proteins, Sema6A in particular, is required for nerve cells in various parts of the brain to migrate to the correct position and make appropriate connections. The Sema6A protein is expressed on the surface of certain cells and is detected by a receptor protein called PlexinA2 on the surface of other cells. If either of these proteins is mutated then a variety of nerve tracts in the brain are miswired. A related protein Sema4D, signals through cells harbouring the receptor protein PlexinB1, which as well as being involved in the development of the nervous system also plays a role in cancer. Tumour progression and metastasis depend on the ability of cancer cells to ensure a blood supply for the delivery of oxygen and nutrients, without it tumours would fail to grow. Sema4D expression is a common feature in some of the most prevalent tumours of epithelial origin where it is believed to promote the development of a blood supply to the growing tumour. Sema4D is also involved in the process of invasive growth, another feature of cancer progression.
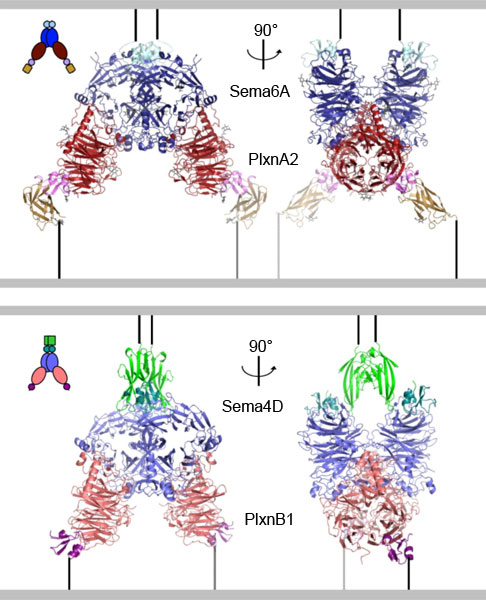
What has not been known until now is exactly how these proteins interact with each other and the effects that this interaction has on the cell. This was finally accomplished using data collected on the macromolecular crystallography beamlines I03 at DIAMOND Light Source and ID23-1 at the ESRF in a collaborative study between researchers at the University of Oxford and Trinity College Dublin. The researchers at Oxford revealed precisely which atoms in the semaphorin, on one cell, interact with which atoms in the plexin on another cell (Figure 1). While results from the researchers in Dublin showed that when the Sema6A-PlexinA2 interaction occurs, the migration of the nerve cells expressing PlexinA2 is reduced. This fits with the known functions of Sema6A, which functions to repel nerve cells or axons from particular areas of the brain. Such atomic resolution ‘snapshots’ of the interaction for two examples of semaphorin and plexin proteins also suggests that this mechanism is quite general for this family. This data provides a model for the effects of the interactions between various members of these large protein families on everything from nervous system development to heart formation and immune system function. These results may also be crucial in trying to develop new therapeutic agents that can block the unwanted functions of these proteins, such as inhibiting Sema4D in metastatic cancer progression.
 |
|
Figure 1. Structures of two semaphorin-plexin complexes reveal that a common mode of interaction underlies semaphorin-plexin cell-cell signalling. |
Principal publication and authors
B.J.C. Janssen (a), R.A. Robinson (a), F. Pérez-Branguli (b), C.H. Bell (a), K.J. Mitchell (b), C. Siebold (a), E.Y. Jones (a), Structural basis of semaphorin-plexin signaling, Nature 467, 1118–1122 (2010); doi:10.1038/nature09468.
(a) Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford (UK)
(b) Smurfit Institute of Genetics and Institute of Neuroscience, Trinity College Dublin (Ireland)
Top image: To be active, the semaphorin-plexin complex must contain two semaphorin dimers independently bound to two plexin molecules. A Cos-7 cell collapse assay shows a) non-collapsed cells and b) a typical collapse seen on media addition of semaphorin dimers.



