- Home
- News
- Spotlight on Science
- X-ray holography...
X-ray holography reveals how the local brain tissue environment affects axon morphology
15-02-2021
Experiments using high-resolution X-ray nano-holotomography at beamline ID16A reveal how axon diameters and trajectories are modulated by other white matter structures. This impacts the non-invasive investigation of axon diameter with diffusion MRI and challenges current knowledge of how axons conduct signals.
The structure of axons, the brain’s communication cables, affects how fast signals are conducted in the brain network. Axon diameter (AD) determines the conduction velocity (CV) of signals along myelinated axons and acts as an indicator of brain performance. It is also a potential biomarker of some neurodegenerative diseases, such as Multiple Sclerosis, that selectively damage certain sizes of axons. Probing the ADs in the living brain would therefore shed light on the health and function of the brain network. Currently, diffusion Magnetic Resonance Imaging (MRI) is the only method capable of doing so, but validation studies show that diffusion MRI overestimates AD when compared to light microscopy. The diffusion MRI approach assumes that axons have very simple cylindrical shapes. To improve the diffusion MRI estimates of AD, therefore, knowledge of the true three-dimensional axon morphologies – beyond that which is possible to obtain with two-dimensional microscopy techniques – is needed [1].
To investigate axon morphology, tissue belonging to the white matter (WM) of a monkey brain was imaged at beamline ID16A using X-ray nano-holotomography (XNH) with a 75-nm isotropic voxel size. The primary sample studied was from the monkey corpus callosum – an organised WM region consisting of many axons aligned in the same direction. The ADs in the corpus callosum of the same monkey brain had previously been studied with diffusion MRI [2,3]. By acquiring four consecutive and overlapping holograms, the cylindrical field-of-view (FOV) spanned a diameter of 153.6 µm and height of 584.5 µm.
Cells, blood vessels and vacuoles could be segmented and quantified within this volume (Figure 1a), demonstrating the intricate architecture of WM in which the cell clusters aligned either with the main direction of the axons, or were anchored to the blood vessels. Axons were segmented from one of the XNH volumes and were shown to have inherently complex morphologies. A diameter analysis concluded that the diameters of axons vary considerably along their lengths (Figure 1b), with larger axons having less specific mean diameters than smaller ones. In other words, axons cannot be described as having well-defined diameters, contrary to what has long been believed. This implies that single axon diameters cannot be accurately estimated from two-dimensional images, such as those from light or electron microscopy. By also quantifying the thickness of the signal-boosting myelin sheath, it could be shown that the diameter changes of axons impact the conduction velocities along them, challenging current knowledge of how axons communicate signals.
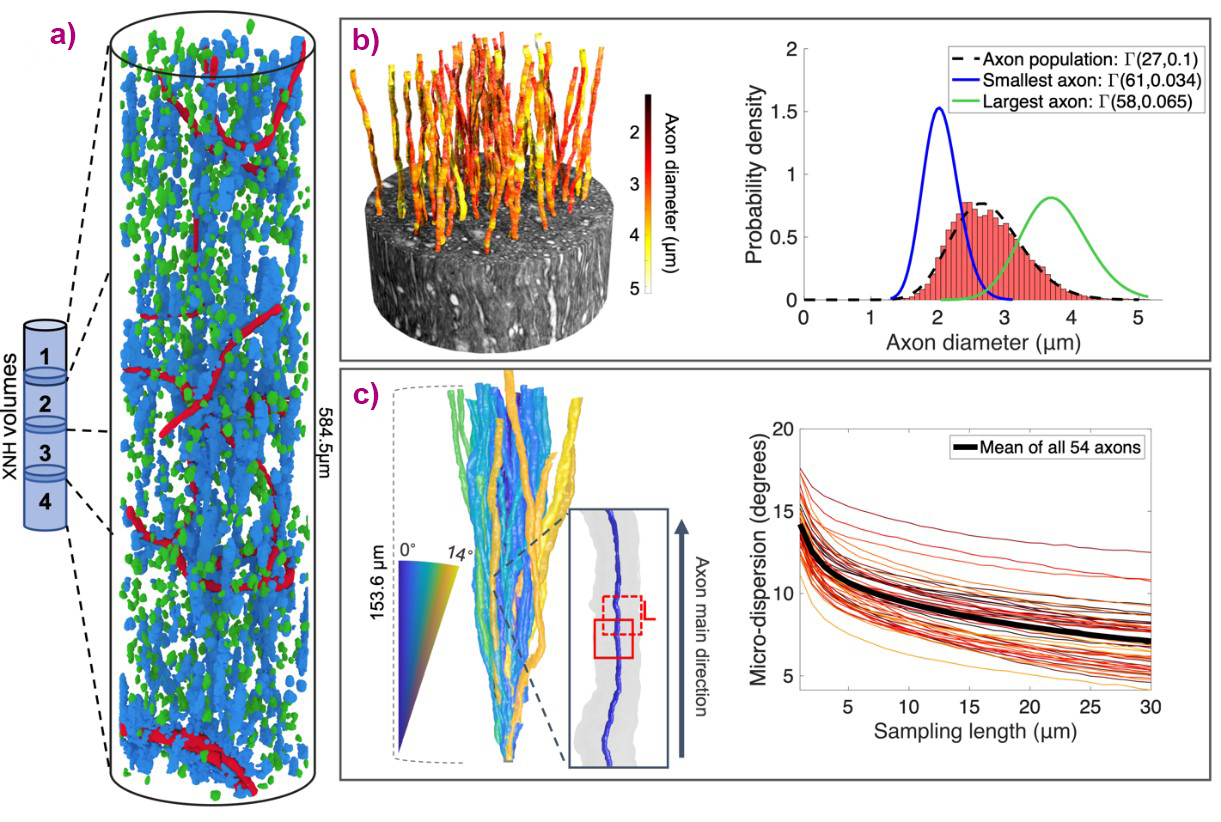
Fig. 1: a) Three-dimensional visualisation of cell cluster (blue), blood vessel (red), and vacuole (green) segmentations in the extended FOV produced by overlapping XNH volumes. b) Left: Segmented axons from the monkey splenium. Right: The total axon diameter distribution (black). The diameter distributions along the thinnest (blue) and thickest (green) axons are also shown. c) The trajectory variations of the axons were quantified in terms of the angle of segments of length L from the main axon direction.
Simulations of diffusion within the axons established how the variations in diameter and trajectory (Figure 1c) of the axons cause an overestimation of AD with diffusion MRI, such as that observed in validation studies. Looking ahead, the geometrical characterisations of the axons obtained from the XNH volumes could act as axonal ‘fingerprints’ to guide the development and validation of diffusion MRI methods for more accurate in-vivo AD estimation.
By studying the morphological variations of axons against the backdrop of the WM environment in Figure 1a, several trends could be exposed. Large structures such as blood vessels were associated with local trajectory changes of the axons, visible almost as a warping effect (Figure 2a). Similarly, axons were shown to skirt around cell clusters (Figure 2b), while diameter decreases could be associated with the presence of vacuoles (Figure 2c). The analysis of an XNH volume from a crossing fibre region of the monkey brain showed how the axons also adapt to each other’s trajectories, sometimes even twisting and spiralling (Figure 2d).
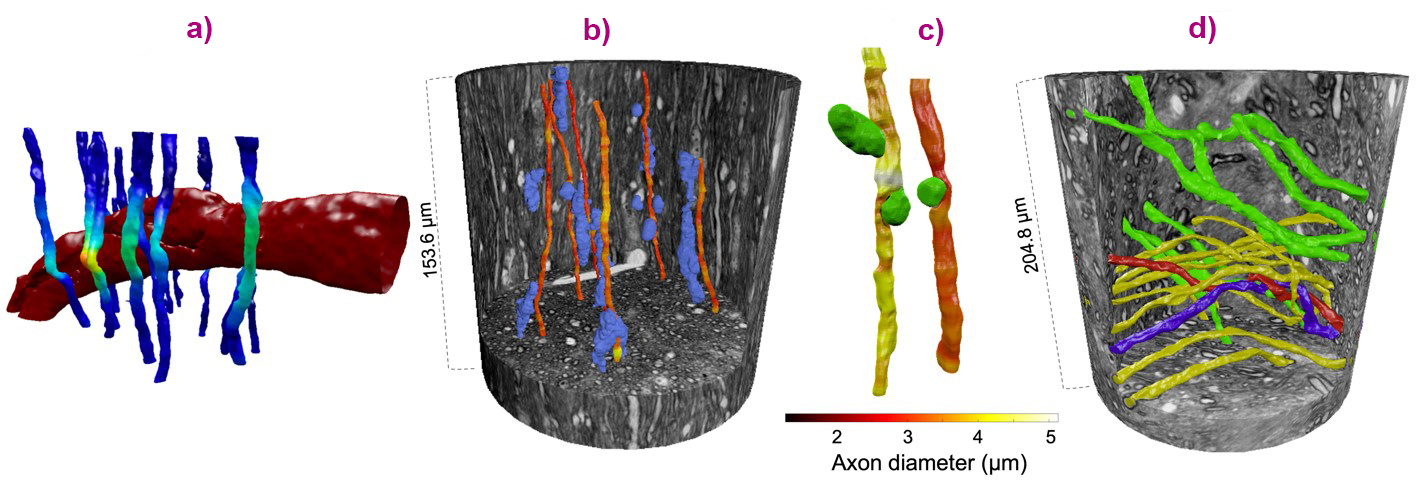
Fig. 2: a) Blood vessels and (b) cell clusters impacted the trajectories of the axons. c) Vacuoles were associated with decreases in diameter. d) In a region of fibre crossings with axons projecting in different directions (yellow vs. green), axons were seen to twist around each other (red and blue).
The discovery that axon morphology can be modulated by extra-axonal structures offers an exciting new avenue for further research. It suggests that axon morphologies may differ in situations where the extra-axonal environment is expected to change, such as in disease. Whether these changes affect the signal conduction process, or if they can be quantified with diffusion MRI, are important research questions to pursue.
Principal publication and authors
Axon morphology is modulated by the local environment and impacts the noninvasive investigation of its structure–function relationship, M. Andersson (a,b), H.M. Kjer (a,b), J. Rafael-Patino (c), A. Pacureanu (d), B. Pakkenberg (e), J-P. Thiran (c,f,g), M. Ptito (h,i), M. Bech (j), A. Bjorholm Dahl (b), V. Andersen Dahl (b), T.B. Dyrby (a,b). Proc. Natl. Acad. Sci. USA 117(52) 33649-33659 (2020); https://doi.org/10.1073/pnas.2012533117.
(a) Danish Research Centre for Magnetic Resonance, Center for Functional and Diagnostic Imaging and Research, Copenhagen University Hospital Hvidovre (Denmark)
(b) Department of Applied Mathematics and Computer Science, Technical University of Denmark, Kongens Lyngby (Denmark)
(c) Signal Processing Laboratory (LTS5), Ecole Polytechnique Fédérale de Lausanne (Switzerland)
(d) ESRF
(e) Research Laboratory for Stereology and Neuroscience, Copenhagen University Hospital, Bispebjerg (Denmark)
(f) Radiology Department, Centre Hospitalier Universitaire Vaudois and University of Lausanne (Switzerland)
(g) Center for Biomedical Imaging (CIBM), Lausanne (Switzerland)
(h) School of Optometry, University of Montreal (Canada)
(i) Department of Neuroscience, Faculty of Health Science, University of Copenhagen (Denmark)
(j) Department of Medical Radiation Physics, Clinical Science, Lund University (Sweden)
References
[1] T.B. Dyrby et al., NeuroImage 182, 62-79 (2018).
[2] D.C. Alexander et al., NeuroImage 52, 1374-1389 (2010).
[3] T.B. Dyrby et al., Magn. Reson. Med. 70, 711-721 (2013).



