- Home
- News
- Spotlight on Science
- Modulation in the...
Modulation in the exit tunnel of the S. aureus ribosome leads to erythromycin resistance
15-01-2020
Ribosomes are strategic targets for antibiotics. Thus, several bacteria have developed antibiotic resistance mechanisms that lead to ribosome mutations. Understanding these mechanisms is crucial for attempts at controlling or minimising antibiotics resistance that is increasing at an alarming rate worldwide.
A different, unforeseen mechanism of resistance to antibiotics has been deciphered structurally for antibiotics that target ribosomes. This is in contrast to the results of previous studies which indicated that most mutations leading to pathogen antibiotic resistance are located in or around the active sites of the ribosomes [1,2]. The unforeseen mechanism was deciphered, using the cryo-electron microscopy station CM01 at the ESRF, where the high-resolution cryo-EM structures of the ribosome of an erythromycin resistant strain of the pathogen S. aureus was determined. This strain harbors a deletion mutation of two amino acids R88-A89 in the β hairpin loop of the ribosomal protein uL22. This mutation leads to erythromycin resistance, although the deletion mutation site is located about 16 Å away from the erythromycin binding site.
Erythromycin is a macrolide antibiotic that targets the ribosome. It is used for the treatment of a number of infections. It acts by binding to the ribosome at the upper rim of the nascent protein exit tunnel (NPET) and blocking the nascent chain progression. It was first isolated in 1952 from the bacteria Saccharopolyspora erythraea. Nowadays erythromycin is mentioned in the World Health Organization's List of Essential Medicines [3], as one of the safest and most effective medicines needed in a health system, but its usage is impaired substantially owing to resistance.
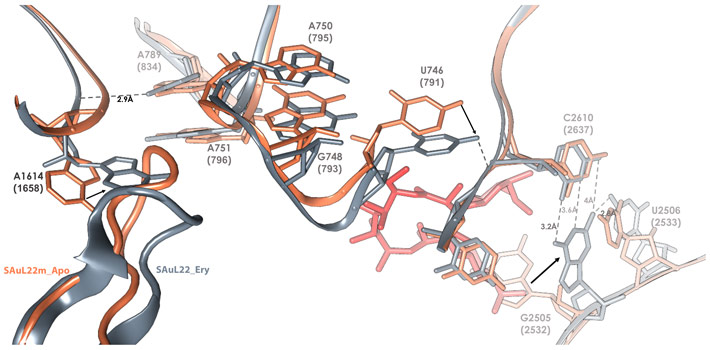
By determining the structures of both the apo (SAuL22m_apo) and erythromycin bound ribosome (SAuL22m_ery) of this mutant, modulation in the shape of the nascent protein exit tunnel (NPET) was identified in the apo structure, which in turn undergoes additional changes upon erythromycin binding (Figure 1). This modulation enables resistance even when erythromycin is bound to the ribosomes.
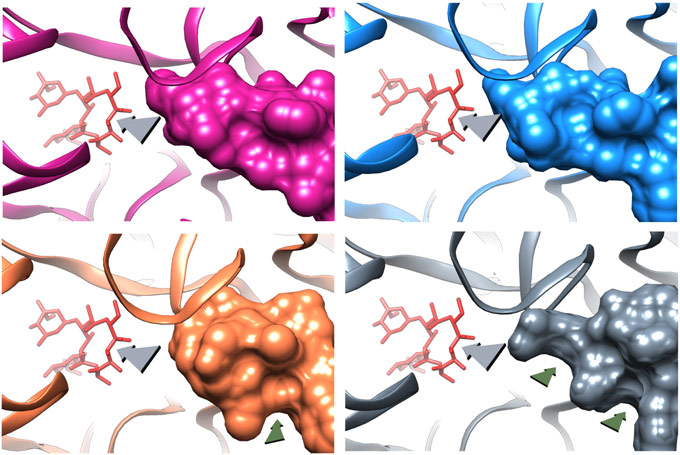
The deletion mutation causes the β hairpin loop of ribosomal protein uL22 to move towards the NPET wall. This movement enables widening of the tunnel by creating two new grooves at the tunnel constriction that is typically formed by the extended loops of the ribosomal proteins uL4 and uL22 protein (Figure 2). Upon erythromycin binding to the mutant ribosome an additional change in the β hairpin loop of ribosomal protein uL22 conformation was detected. These structural adjustments enable the growing nascent protein to bypass erythromycin, and to progress along the tunnel to exit the ribosome.
Deciphering the structural aspects of this resistance mechanism, at high resolution, may lead to the design of erythromycin-based antibiotics that can be effective against such mutations.
Principal publication and authors
Exit tunnel modulation as resistance mechanism of S. aureus erythromycin resistant mutant, Y. Halfon (a), D. Matzov (a), Z. Eyal (a), A. Bashan (a), E. Zimmerman (a), J. Kjeldgaard (b), H. ingmer (c) & A. Yonath (a), Scientific Reports 9, 11460 (2019); DOI: 10.1038/s41598-019-48019-1.
(a) The Weizmann Institute of Science, The Department of structural biology, Rehovot (Israel)
(b) National Food Institute, Technical University of Denmark, Lyngby (Denmark)
(c) Department of Veterinary and Animal Sciences, Faculty of Health and Medical Sciences, University of Copenhagen, Frederiksberg (Denmark)
References
[1] Ribosomal antibiotics: contemporary challenges, T. Auerbach-Nevo, D. Baram, A. Bashan, M. Belousoff, E. Breiner, C. Davidovich, A. Yonath, Antibiotics (Basel) 5, 24 (2016); doi: 10.3390/antibiotics5030024.
[2] Structural insights of lincosamides targeting the ribosome of Staphylococcus aureus, D. Matzov, Z. Eyal, R.I. Benhamou, M. Shalev-Benami, Y. Halfon, M. Krupkin, A. Yonath, Nucleic Acids Res 45, 10284-10292 (2017). doi: 10.1093/nar/gkx658.
[3] World Health Organization. Executive summary: the selection and use of essential medicines 2019: report of the 22nd WHO Expert Committee on the selection and use of essential medicines: WHO Headquarters, Geneva, 1-5 April 2019.