- Home
- News
- Spotlight on Science
- Condensin complexes...
Condensin complexes fasten the belt around DNA
20-03-2018
The crystal structure has been determined of the major DNA binding site in the eukaryotic condensin complex, which is jointly formed by the Cnd3 HEAT-repeat and Cnd2 kleisin subunits. The DNA-bound structure indicates an unconventional ‘safety belt’ mechanism that anchors condensin complexes and might provide condensin with the means to form large chromatin loops.
Ever since the first observation of condensed chromosomes in dividing cells during the late 19th century, scientists have been fascinated by trying to understand the principles that underlie this dramatic reorganisation of chromatin fibres [1]. Since then, it took more than one hundred years until the five-subunit condensin complex was identified as the leading player in the formation of mitotic chromosomes [2]. Despite its key role for the correct segregation of chromosomes during mitosis and meiosis, the molecular mechanisms by which condensin complexes interact with chromatin fibres and alter their topology have so far remained largely mysterious.
Condensin complexes share their basic architecture with other structural maintenance of chromosomes (SMC) complexes, including cohesin and SMC5/6 in eukaryotes, and different SMC-type complexes in prokaryotes. All SMC complexes play essential roles in the maintenance of a cell’s genetic material during various aspects of chromosome biology, including chromosome condensation and segregation, DNA recombination and damage repair, and the organisation of chromosomal territories. The conserved basic architecture of these complexes suggests that they share a common mode of action in how they perform this array of seemingly diverse tasks.
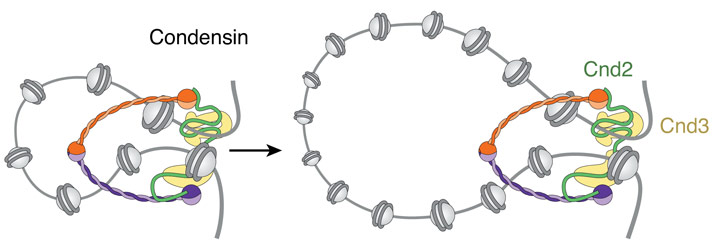
How could such a common mechanism work? A number of different models has been suggested over the years, but recently a process called ‘loop extrusion’ has been championed as a simple and elegant mechanism that could explain most of the cellular tasks of SMC complexes (Figure 1). How these complexes might form and extend DNA loops is, however, not understood, but one precondition of any loop extrusion model is that the proteins would need to make direct contact with their DNA substrates. Since all subunits of SMC complexes are largely devoid of classic DNA-binding motifs, the nature and specificity of this interaction has remained uncertain.
Previous work had demonstrated that a ternary sub-complex consisting of two HEAT repeat subunits and the so-called kleisin subunit can facilitate DNA binding in a sequence-unspecific manner [3]. To identify the DNA binding site in this ternary sub-complex, the three subunits were expressed and purified individually and tested alone or in combination for DNA binding. Neither subunit alone could bind DNA efficiently. Only the combination of the HEAT-repeat protein Cnd3 with the kleisin Cnd2 yielded a DNA-binding activity comparable to the ternary complex.
This complex was – after some additional engineering and screening – amenable to crystallisation by vapour diffusion. Following data collection at beamline ID29, the structures from two different yeast species were determined to 2.6 and 2.8 Å resolution, respectively. The structures not only provided the molecular details of how the kleisin subunit binds along the inner surface of the harp-shaped HEAT-repeat solenoid, but also gave a clear indication of the DNA binding groove in the complex. Mutation of conserved positively charged residues in this groove confirmed its functional importance for recruiting condensin to chromosomes both in vitro and in vivo.
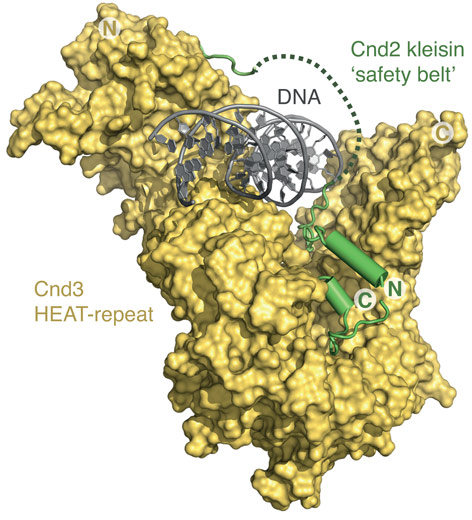
Additional extensive DNA screening yielded co-crystals of the sub-complex bound to DNA, for which data was collected at ID23-2 and determined a 3.0 Å structure by molecular replacement (Figure 2). The resulting structure not only confirmed that the DNA double helix passes through the positively charged groove but furthermore suggested that the kleisin subunit encircles the bound DNA – in a manner reminiscent of the way a safety-belt fastens a seated passenger. Based on in vitro site-specific cross-linking and DNA binding experiments, as well as chromatin co-localization experiments, it was possible to show that this mode of interaction provides a simple and elegant mechanism to tightly bind a DNA substrate without the need for specific interactions with the DNA bases.
 |
|
Figure 2. Structure of the Cnd2–Cnd3 subcomplex bound to double-stranded DNA. Entrapment of the DNA helix by the Cnd2 ‘safety belt’ is indicated by a dotted line (disordered in the crystal). |
Taken together, the results provide molecular insights into the architecture of eukaryotic condensin complexes and identify a major DNA binding site in the complex, formed by its Cnd3 HEAT-repeat and Cnd2 kleisin subunits. The new structures explain the preference for double-stranded DNA binding and lack of sequence specificity. The findings suggest a sophisticated DNA ‘safety belt’ mechanism that can be used to anchor condensin complexes to chromosomes and thereby enables the formation of large chromatin loops as the fundamental organisational principle of higher-order chromatin structure.
Principal publication and authors
Structural Basis for a Safety-Belt Mechanism That Anchors Condensin to Chromosomes, M. Kschonsak (a), F. Merkel (a), S. Bisht (a), J. Metz (a), V. Rybin (b), M. Hassler (a) and C.H. Haering (a), Cell 171, 588-600 (2017); doi: 10.1016/j.cell.2017.09.008.
(a) Cell Biology and Biophysics Unit, Structural and Computational Biology Unit, European Molecular Biology Laboratory (EMBL) Heidelberg (Germany)
(b) Protein Expression and Purification Core Facility, European Molecular Biology Laboratory (EMBL) Heidelberg (Germany)
References
[1] W. Flemming, in Zellsubstanz, Kern und Zelltheilung (1882).
[2] Hirano et al., Cell 89, 511-521 (1997).
[3] Piazza et al., Nat. Struct. Mol. Biol. 21, 560-568 (2014).