- Home
- News
- Spotlight on Science
- 3D structure sheds...
3D structure sheds light on the mechanism of uromodulin self-polymerisation
30-05-2016
Urinary tract infection is the most common non-epidemic bacterial infection in humans. The urinary tract lacks a mucus layer to shield it against infections and mammals produce a molecular net that captures pathogenic bacteria in the urine and clears them from the body. Uromodulin, first isolated more than 60 years ago, has been recognised as a guardian against urinary tract infection and a crucial player in innate immunity. Massively produced in the kidneys, uromodulin self-assembles into long polymers which together with salt ions form a water-soluble net. The three-dimensional structure of the polymerisation region of human uromodulin provides insights into how this important protein forms polymers, and how it is compromised by mutation in patients with kidney diseases.
Urinary tract infection (UTI) is the most common non-epidemic type of bacterial infection: every year there are more than 150 million cases, with a healthcare cost of more than 6 billion dollars. Women are particularly prone to UTI, resulting in more than fifty percent lifetime risk; in men UTI is less common, but - when they occur - they can be very serious. Because the human urinary tract lacks a mucus layer to shield it against infections, our body developed a different protection strategy. Every day, our kidneys produce massive amounts of a sugar-rich molecule known as uromodulin (UMOD) or Tamm-Horsfall protein. UMOD sugar units resemble those of proteins exposed on the urothelial surface and act as a molecular decoy that is mistakenly bound by uropathogenic bacteria trying to invade the urinary tract. The filamentous structure of UMOD thereby neutralises and permits expulsion of the dangerous bacteria upon urination.
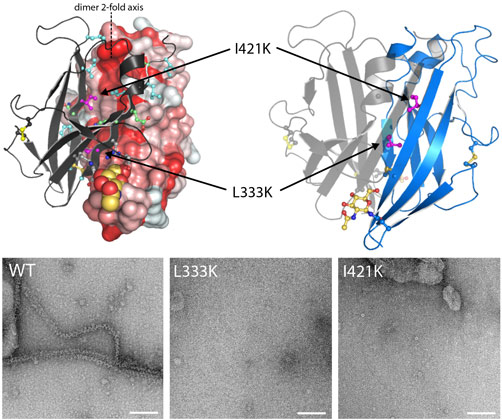
The highly complex structure of UMOD has long hindered its detailed biochemical analysis. By exploiting a novel mammalian cell expression strategy [1], we have successfully produced the whole polymerisation region of human UMOD and determined its 3D structure using X-ray crystallography, with data collected at beamline ID29. This provided a molecular description of how the polymerisation of UMOD is controlled. The electron density map of UMOD reveals that a structurally-ordered linker connects the ZP-N and ZP-C domains that constitute the zona pellucida module of UMOD. Coupling of the linker to ZP-C induces an extended conformation of the ZP module, which in turn exposes the βA/βG surface of ZP-N to form a dimer that initiates homopolymerisation (Figure 1). UMOD ZP-N dimerisation is mediated by an extensive hydrophobic interface, whose formation is required for protein assembly into filaments.
Comparison of the 3D structure of this polymerisation region of UMOD with that of a structurally-related egg coat component, ZP2, has led to an explanation of why UMOD can self-assemble whereas the egg coat component, ZP2, can only form filaments in the presence of sperm receptor ZP3. Most importantly, the atomic architecture of UMOD helps us to understand many patient mutations that are associated with kidney disease, as well as non-syndromic deafness-causing mutations in another structurally-related but functionally-distinct molecule, inner ear protein α-tectorin. Finally, the structure of UMOD constitutes a valuable framework to help us understand the biology of pancreatic and M-cell glycoprotein 2 (GP2), a highly similar homopolymeric molecule that has been recently recognised as the major target of the antipancreatic antibodies found in Crohn’s disease.
Principal publication and authors
A structured interdomain linker directs self-polymerization of human uromodulin, M. Bokhove (a), K. Nishimura (a), M. Brunati (b), L. Han (a), D. de Sanctis (c), L. Rampoldi (b), L. Jovine (a), Proc Natl Acad Sci U S A. 113, 1552-7 (2016); doi: 10.1073/pnas.1519803113.
(a) Department of Biosciences and Nutrition & Center for Innovative Medicine, Karolinska Institutet, Huddinge (Sweden)
(b) Molecular Genetics of Renal Disorders Unit, Division of Genetics and Cell Biology, San Raffaele Scientific Institute, Milan (Italy)
(c) ESRF
References
[1] M. Bokhove, H. Sadat Al Hosseini, T. Saito, E. Dioguardi, K. Gegenschatz-Schmid, K. Nishimura, I. Raj, D. de Sanctis, L. Han, L. Jovine, J Struct Biol. 194, 1-7 (2016).
Top image: Self-polymerisation of uromodulin.