- Home
- News
- Spotlight on Science
- 3D imaging of whole...
3D imaging of whole cells using cryocooled coherent X-ray diffraction
26-10-2015
X-ray imaging of whole cells using coherent diffraction has the potential to bridge the gap between optical and electron microscopy. However, radiation damage has proven limiting. An international team of scientists, including the ESRF, have now demonstrated that cryocooling can be used in conjunction with coherent diffractive imaging (cryo-CDI) to reveal the 3D structure of a parasitic cell with an overall spatial resolution better than 100 nm.
A long way from Robert Hooke’s first description of cells in cork, we are now experiencing a revolution in cellular imaging. Optical microscopes are peering more closely into cells than ever before, revealing their inner workings in real time and with high spatial resolution. Thanks to technical and computational improvements, electron microscopy now approaches atomic resolution in images of single protein molecules. Bridging the gap between these two imaging modalities is X-ray microscopy. Coherent X-ray microscopy is a young technique that seeks to capitalise on recent improvements in synchrotron science that have resulted in in brighter and more coherent X-ray beams. It makes use of a technique known as coherent X-ray diffraction imaging (CDI) [1] to recover a quantitative map of the 3D structure of a cell.
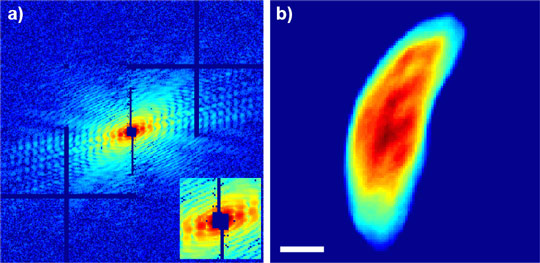
We used a method similar to that employed by computed tomography (CT). The process involves a synthesis of multiple projections of the same object from different angles. The difference between CT and CDI is that CT relies on projections, which are transmission images of an object, while CDI interprets the complex pattern of X-rays diffracted from the object as it is illuminated by a coherent X-ray beam (Figure 1). CDI is constrained by what is commonly known as the phase problem in X-ray diffraction. That is, half of the data required to produce an interpretable image of the object is missing. By using a lens to collect diffracted light, optical microscopes overcome the phase problem. To solve this problem for X-rays, CDI instead tasks powerful computer algorithms with synthesis and interpretation of the diffracted data. In effect, the computer replaces an X-ray lens. This process guarantees the most efficient detection of the diffracted X-rays and a faithful recovery of the object structure.
J. Miao, who leads the present study, first experimentally demonstrated CDI in 1999 [2]. Since that initial demonstration, CDI has been used to image a broad range of samples. These include nanoscale objects such as inorganic nanoparticles and even single viruses. Larger, micron-scale objects such as whole cells have also been imaged. Biological specimens pose a particular challenge for CDI since they scatter X-rays weakly. While higher X-ray doses can compensate for such weak scattering, cells and other biomaterials are sensitive to radiation damage incurred by exposure to high doses of X-rays. To ensure sample integrity, the X-ray doses used to inspect the cell must be kept at a minimum.
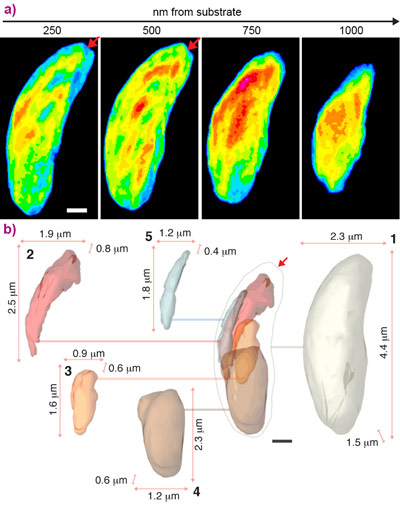
Cryogenic preservation techniques can be used to safeguard the integrity of a cell during exposure to high doses of X-rays. By rapidly freezing the cell while in its native environment, cryogenic techniques maintain that cell in a deeply frozen state that allows its long-term exposure to X-rays. In a proof-of-principle experiment, we combined cryogenic techniques and 3D CDI at beamline ID10 to reveal the architecture of a cell at high spatial resolution. In this particular example, we investigated a parasite known as Neospora caninum. This species of cell primarily infects dogs and causes abortions in livestock. While it does not infect humans, it is closely related to Plasmodium and Toxoplasma, the causative agents of malaria and toxoplasmosis. The distinctive banana shape of the cell and the organisation of its inner structures are important for infection of its hosts. These features are highlighted in the recovered structure (Figure 2).
The 3D imaging of a Neospora caninum with an overall spatial resolution better than 100 nm represents the first experimental demonstration of cryo-CDI for quantitative 3D imaging of whole, frozen-hydrated cells using hard (8 keV) X-rays. As new, brighter, and highly coherent X-ray sources continue to emerge worldwide, our work forecasts the routine and quantitative 3D imaging of whole cells with spatial resolutions in the tens of nanometres. These studies are important building blocks for the better structural understanding of cells, and may lead to advances in the fields of imaging, biology and medicine, as well as the general realm of nano and biomaterials.
Principal publication and authors
Three-dimensional coherent X-ray diffractive imaging of whole frozen-hydrated cells, J.A. Rodriguez (a), R. Xu (b), C.-C. Chen (c), Z. Huang (d), H. Jiang (e), A.L. Chen (f), K.S. Raines (g), A. Pryor Jr (b), D. Nam (h), L. Wiegart (i), C. Song (h), A. Madsen (j), Y. Chushkin (k), F. Zontone (k), P.J. Bradley (f) and J. Miao (b), IUCrJ 2, 575–583 (2015); doi: 10.1107/S205225251501235X.
(a) UCLA-DOE Institute for Genomics and Proteomics, University of California, Los Angeles (USA)
(b) Department of Physics and Astronomy and California NanoSystems Institute, University of California (USA)
(c) National Sun Yat-sen University, Kaohsiung (Taiwan)
(d) Carl ZEISS X-ray Microscopy Inc., Pleasanton (USA)
(e) Shandong University, Jinan (People's Republic of China)
(f) Department of Microbiology, Immunology, and Molecular Genetics, University of California, Los Angeles (USA)
(g) Department of Applied Physics, Stanford University (USA)
(h) Department of Physics, Pohang University of Science and Technology (South Korea)
(i) NSLS-II Photon Sciences Division, Brookhaven National Laboratory, Upton (USA)
(j) European X-ray Free Electron Laser, Hamburg (Germany)
(k) ESRF
References
[1] J. Miao, T. Ishikawa, I.K. Robinson, & M.M. Murnane, Science 348, 530–535 (2015).
[2] J. Miao, P. Charalambous, J. Kirz, & D. Sayre, Nature 400, 342–344 (1999).
Top image: A 2D projection of a reconstructed 3D Neospora caninum cell. Scale bar: 500 nm.