- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2013
- Structural biology
- Long wavelength S-SAD phasing of the crystal structure of FAM3 PANDER reveals a new class of signalling molecules
Long wavelength S-SAD phasing of the crystal structure of FAM3 PANDER reveals a new class of signalling molecules
Cytokines are a diverse group of soluble proteins or peptides that act as signalling molecules in the body [1]. FAM3B pancreatic derived factor (PANDER) was initially suggested to be a cytokine [2] is involved in insulin signalling and thereby a potential drug target for diabetes. The FAM3 superfamily was predicted to contain classical four-helix bundle cytokines, featuring a typical up-up-down-down fold.
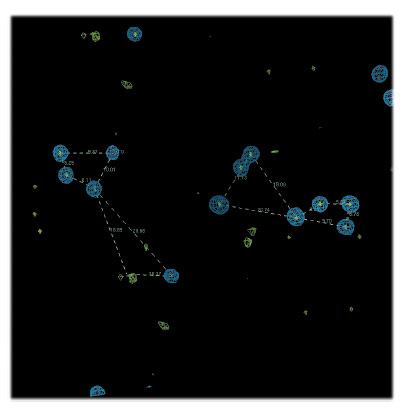
Despite the success of sulfur SAD phasing for high-resolution systems, novel structures solved from low symmetry or moderate resolution data are still rare. Here we present the S-SAD phasing of FAM3 PANDER. To optimise the PANDER sulfur SAD signal-to-noise ratio, highly redundant data were collected at beamline ID29 in two kappa wedges using a photon energy of 6 keV (λ ≈ 2.06 Å). Due to the moderate resolution, the anomalous signal did not extend beyond 4.5 Å and an extensive substructure search did not result in any obvious hits. However, by careful analysis and crosschecking of the solutions, a number of recurring super-sulfur atoms could be assigned and split into separate atoms (Figure 62). Density modification and NCS averaging identified the remaining sulfur sites, extending the phases to about 3.7 Å resolution. To maximise the phasing power, the substructure parameters were iteratively re-refined using density modified Hendrickson Lattman coefficients as phase restraints, gradually increasing the solvent content. The new phase-estimates were finally subjected to statistical density modification and the phases extended to 2.4 Å resolution.
 |
|
Fig. 62: NCS-relation between the two PANDER molecules in the asymmetric unit of the crystals studied. Partial S-substructure in blue, anomalous LLG map in green. |
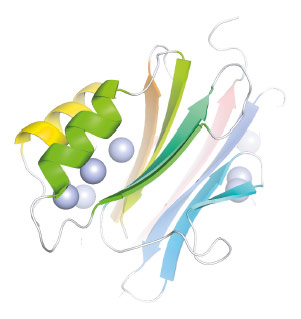
Even from initial low resolution maps it was clear that PANDER is not an all-α helical protein and, contrary to the previous suggestions, PANDER exhibits a novel globular β-β-α fold (Figure 63). The structure is composed of two antiparallel β sheets lined by three short helices. This fold shares no relation to the predicted cytokines but is clearly conserved throughout the FAM3 superfamily.
 |
|
Fig. 63: The FAM3 PANDER β-β-α-fold coloured from blue to red N to C terminus. Sulfur atoms indicated by spheres. |
A search using the PANDER structure as a template in the DALI fold database resulted in a small number of hits with remotely related secondary structure. The most similar structure, with a Z-score of 5.1, was the artificially designed TOP7 protein (PDB code 1QYS), sharing a related C-terminal β-α architecture. The lack of naturally occurring structures with related topology and the partial similarity to TOP7, designed to explore new regions of the protein universe [3] indicates that PANDER represents a novel protein fold.
Since the discovery of FAM3B, the view of its function has broadened and become more complex: from an islet specific cytokine with apoptotic activity to that of a hormone involved in negative regulation of beta cell function and liver glucose homeostasis. However, down-stream signalling mechanism(s) for PANDER remains to be elucidated including the identification of a PANDER-specific receptor.
The unexpected PANDER fold together with the potent biological actions of proteins in the superfamily indicate that FAM3 might represent a new class of signalling molecules, strikingly different from the known four-helix bundle cytokines.
Principal publication and authors
P. Johansson (a), J. Bernström (b), T. Gorman (c), L. Öster (a), S. Bäckström (b), F. Schweikart (b), B. Xu (b), Y. Xue (a) and L. Holmberg Schiavone (b), Structure 21, 306–313 (2013).
(a) Structure and Biophysics, Discovery Sciences, AstraZeneca, Mölndal (Sweden)
(b) Reagents and Assay Development, Discovery Sciences, AstraZeneca, Mölndal (Sweden)
(c) High-Content Biology, Discovery Sciences, AstraZeneca, Alderley Park, Macclesfield (UK)
References
[1] X. Wang, P. Lupardus, S.L. Laporte and K.C. Garcia, Annu Rev Immunol. 27, 29-60 (2009).
[2] Y. Zhu, G. Xu, A. Patel, M.M. McLaughlin, C. Silverman, K. Knecht, S. Sweitzer, X. Li, P. McDonnell and R. Mirabile et al. Genomics 80, 144-150 (2002).
[3] B. Kuhlman, G. Dantas, G.C. Ireton, G. Varani, B.L. Stoddard and D. Baker, Science 302, 1364–1368 (2003).



