Highlights
How a therapeutic antibody helps the body to kill tumours
Authors: Na, Z1., Yeo, S. P2., Bharath, S. R.1, Bowler, M.W.3, Balikci, E.1, Wang, C-I2. and Song, H1,4.
- Institute of Molecular and Cellular Biology, Singapore
- Singapore Immunology Network, Singapore
- European Molecular Biology Laboratory, Grenoble Outstation, France
- Department of Biochemistry, National University of Singapore, Singapore
One of the fastest growing areas of novel pharmaceutical treatments is therapeutic antibodies, where the body’s immune system can be stimulated to remove specific cells or proteins. This type of treatment has been hugely successful in the treatment of cancer but often very little is known about the molecular interactions involved.
One such therapeutic antibody is called Pembrolizumab that has been shown to be highly effective in the treatment of melanoma, lung cancer and head and neck carcinoma. The drug works by binding to a cell surface receptor called programmed cell death 1 (PD-1). PD-1 is an immune inhibitory transmembrane receptor that is an immunity checkpoint that prevents the body destroying its own cells by recognising another protein called PD-L1. However, this protein is overexpressed in many types of tumour, allowing the cancer to escape destruction by the immune system.
Using the fully automated service on MASSIF-1 several hundred crystals of a complex between the human receptor PD-1 and Pembrolizumab were screened. The complex is large and flexible leading to a large variation in the quality of crystals. The automation on MASSIF-1 allowed a large number of crystals to be screened rapidly and data collection algorithms managed to collect an optimised data set to 2.9 Å from one crystal. The structure revealed how Pembrolizumab binds to PD-1 and competes with PD-L1. The results should lead to specifically designed therapeutic antibodies with improved efficacy in treating cancer.
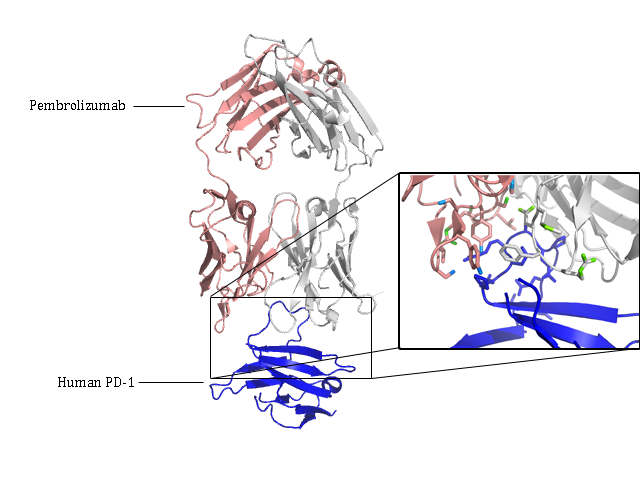
The structure of Pembrolizumab (light chain pink, heavy chain grey) bound to human cell surface receptor PD-1(blue. The molecular details of the interaction are shown in detail in the zoom.
[Na, Z., Yeo, S.P., Bharath, S.R., Bowler, M,W., Balikci, E., Wang, C-I. and Song, H.W. Structural basis for blocking PD-1-mediated immune suppression by therapeutic antibody pembrolizumab (2017) Cell Res. 27, 147-150]
The structure of Rpf2-Rrs1 explains its role in ribosome biogenesis
Authors: Satyavati Kharde, Fabiola R. Calvino, Andrea Gumiero, Klemens Wild and Irmgard Sinning
Heidelberg University Biochemistry Center (BZH), INF 328, D-69120 Heidelberg, Germany
Ribosomes are nano-machines that translate the genetic information encoded by messenger RNA into proteins in all living cells. They are themselves composed of both proteins and RNA with up to 50 separate subunits. Their biogenesis is therefore a complex and highly regulated process that requires many separate assembly factors. The Rpf2-Rrs1 complex is known to be an assembly factor important in the final stages of ribosome formation but until recently its structure and its mode of action was unknown.
Using the automatic and flexible service offered by MASSIF-1, Kharde et al. determined the crystal structure of the complex de novo. Initially, 70 crystals of the native complex were obtained and rapidly screened on MASSIF-1. This resulted in 10 data sets from crystal forms with different space groups and resolution limits. As molecular replacement failed to produce a successful structure solution a second session was booked with 40 crystals of the selenomethionine derivative of the Rpf2-Rrs1 complex screened. This resulted in 10 data sets with the best diffraction to 1.6 Å resolution. These data sets were processed automatically resulting in 9 of the datasets being phased by ESRF automatic SAD protocols. The automatic service on MASSIF-1 allowed the researchers to screen a large number of crystals when they became available, through flexible booking of beamtime, and rapidly feed results back into the project to improve crystal quality and phase the structure by single wavelength anomalous diffraction within 5 weeks.
The high resolution model obtained on MASSIF-1 was then docked into Cryo-electron microscopy density of the pre-formed ribosome. The combination of low and high resolution models has provided an explanation for how this complex recruits the final protein subunits into the machine.
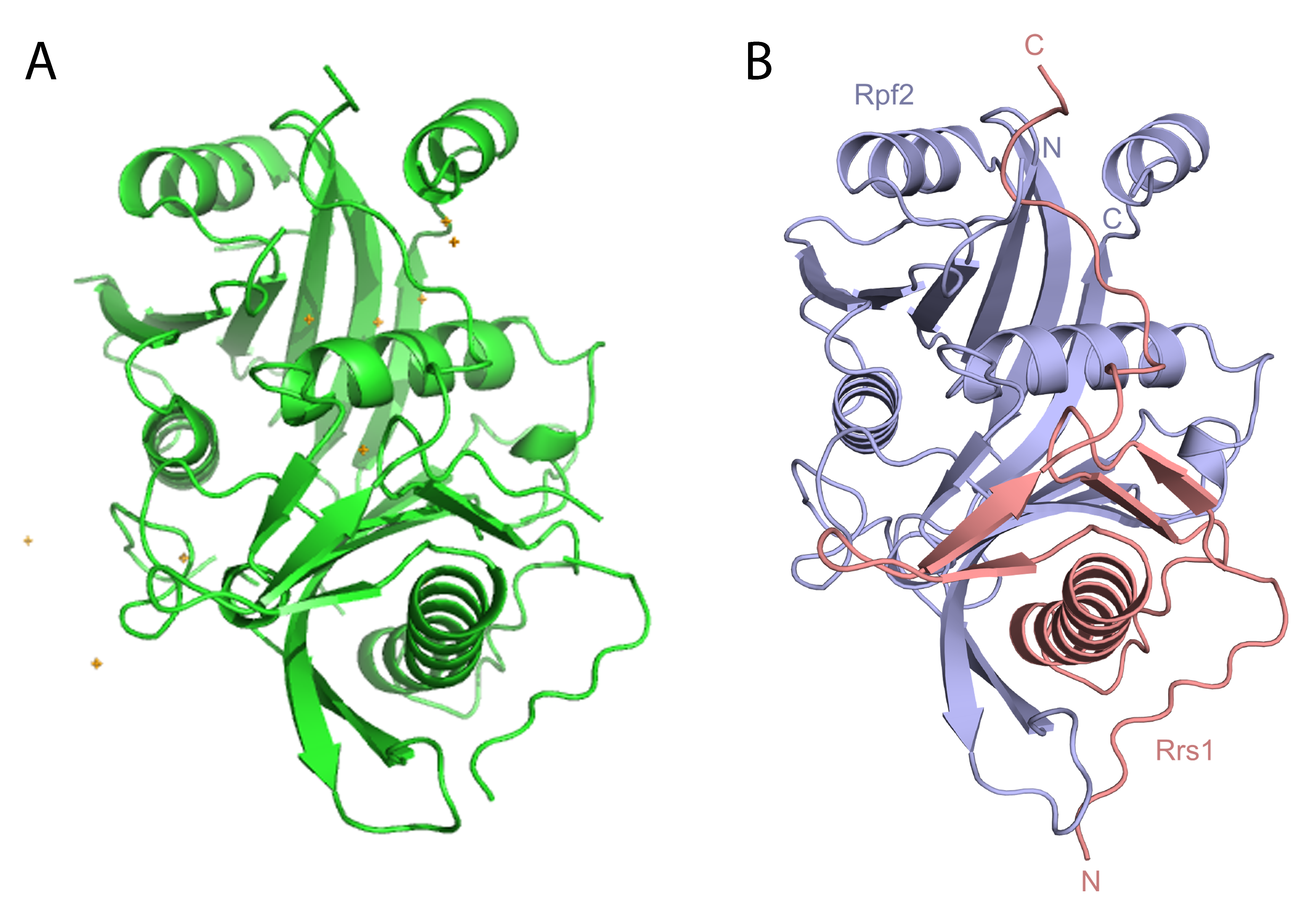
The Rpf2-Rrs1 complex structure solved using diffraction data collected fully automatically on MASSIF-1. A. The model determined by the automatic SAD pipeline is shown as a cartoon from the Cα trace with the Se atom substructure shown as gold crosses. B The final refined model is shown with Rpf2 in magenta and Rrs1 in salmon.
[Kharde, S., Calvino, F. R., Gumiero, A., Wild, K. & Sinning, I. (2015). The structure of Rpf2-Rrs1 explains its role in ribosome biogenesis, Nucleic Acids Res. 43, 7083-7095]



