- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2008
- Soft condensed matter
- DNA adsorption at liquid/solid interfaces studied by X-ray reflectivity
DNA adsorption at liquid/solid interfaces studied by X-ray reflectivity
DNA interactions with surfaces can be repulsive, leading to confinement or depletion, or attractive. A repulsive interaction exists between negatively charged phosphates of DNA and phospholipids of membranes and contributes to enclose the genetic material within the cell. Depletion forces also confine DNA within viral capsids. Repulsive interactions are a common requirement in micro- and nanofluidics. Attractive interactions lead to adsorption. Chemisorption is at the core of solid-phase synthesis of polynucleotides. Physisorption is also of wide technological interest and biological relevance. In hybridisation techniques such as Southern blots, northern blots, or biochips, nucleic acids are immobilised on surfaces, either through chemisorption or physisorption. Chromatographic techniques rely on a reversible immobilisation of DNA. It is thus of great importance to understand the details of DNA interactions with interfaces, liquid/solid, liquid/liquid [1] or vapor/liquid [2].
 |
|
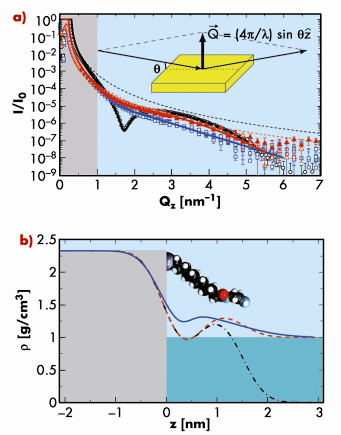
Fig. 47: Synchrotron X-ray reflectivity a) measurements and b) density profile model. |
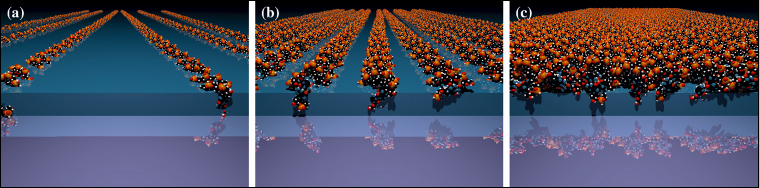
Few experimental techniques can be employed to study the conformation of biological macromolecules on a molecular length-scale. Atomic force microscopy, for example, can be carried out within a liquid solution and has yielded impressive atomic-resolution images of strongly adsorbed double-stranded DNA molecules on a solid substrate. Here we performed X-ray reflectivity studies at the aqueous buffer/solid substrate interface to detect the adsorption of a monomolecular layer of DNA molecules onto (and into) a soft, organic functionalisation layer, positively charged, covalently grafted on an oxide-free, atomically flat, single-crystal silicon wafer. Experiments were carried-out on beamline ID10B using 22.2 keV X-rays to traverse a 30 mm water path-length. The originality of the method was to characterise the grafted substrates at two different density contrasts (Figure 47): measured first under air, then under buffer solution, and then to analyse the data conjointly, as is often the practice for neutron scattering. This severely constrained the model parameters yielding a simple, robust measurement of the soft structure. Thus, the very subtle change observed in situ upon adsorption of the somewhat disordered monomolecular layer of DNA could be reliably interpreted in terms of adsorbed mass and therefore quantified. The high density of this layer then implies that the thin layer of monodisperse DNA double-stranded chains must exhibit some 2D nematic ordering (Figure 48), although this is not observable directly in the experiment.
 |
|
Fig. 48: Molecular models of the organisation of the adsorbed DNA layer a) full embedding, b) intermediate and c) full coverage. |
This interpretation of the results goes against common beliefs about strong adsorption of macromolecules. This adsorption can be irreversible, yet the molecules still retain some mobility (rolling or sliding) to allow rearrangements and compaction through annealing. Even strongly adsorbed on a solid surface, DNA molecules remain accessible to molecular interactions with enzymes, for example, as can be shown through some elegant biochemical experiments [3]. The tethering of single molecules extending into the free solution is therefore not strictly necessary for biotechnological applications and this opens possibilities for the conception of new hybridisation biochips, in particular, with aims of taking advantage of electrical coupling with the oxide-free semiconductor substrates. Enhanced efficiency of the hybridisation reaction can also be gained through a reduction of dimensionality [4] on adsorption of the reactants to a surface.
Principal publications and authors
C. Douarche (a,b,c), R. Cortès (a), S.J. Roser (d), J.-L. Sikorav (e) and A. Braslau (f), J. Phys. Chem. B 112, 13676–13679 (2008); C. Douarche (a,b), R. Cortès (a), C. Henry de Villeneuve (a), S.J. Roser (c), and A. Braslau (e), J. Chem. Phys. 128, 225108 (2008).
(a) Physique de la Matière Condensée, École Polytechnique, CNRS, Palaiseau (France)
(b) Institut de Recherche Interdisciplinaire, Villeneuve d’Ascq (France)
(c) Service de Biologie Intégrative et de Génétique Moléculaire, CEA/Saclay, Gif-sur-Yvette (France)
(d) Department of Chemistry, University of Bath (UK)
(e) Institut de Physique Théorique, CNRS URA 2306, CEA/Saclay, Gif-sur-Yvette (France)
(f) Service de Physique de l’État Condensé, CNRS URA 2464, CEA/Saclay, Gif-sur-Yvette (France)
References
[1] A. Goldar and J.-L. Sikorav, Eur. Phys. J. E 14, 211 (2004).
[2] C. Douarche, J.-L. Sikorav, and A. Goldar, Biophys. J. 94, 134 (2008).
[3] D. Rhodes and A. Klug, Nature (London) 286, 573 (1980).
[4] G. Adam and M. Delbrück, Reduction of Dimensionality in Biological Diffusion Processes. In Structural Chemistry and Molecular Biology; A. Rich and N. Davidson, Eds.; W. H. Freeman and Company: San Francisco, 198 (1968).



