- Home
- News
- Spotlight on Science
- Surface glycoprotein...
Surface glycoprotein coat structure helps explain how trypanosomes avoid detection in their host
16-01-2018
The variant surface glycoproteins (VSGs) of Trypanosoma brucei are essential for survival of the unicellular parasite in its mammalian host. Here, the first complete structures of two VSGs were determined, which give new insights into the properties of a highly dynamic protein shield on the cell surface.
It has long been established that African trypanosomes possess a dense protein surface coat that must offer protection from destruction by the host immune system. The coat is comprised of close to only one GPI-anchored surface protein from a large family of variant surface glycoproteins (VSGs). This VSG is thought to form an impervious, physical barrier on the cell surface, thus protecting the underlying proteins from immune attack. It is itself highly immunogenic, but bound antibodies are passively cleared from the cell surface by hydrodynamic drag forces. Importantly, the expressed VSG can be replaced by another, antigenically distinct, member of the large family of VSGs in a process known as antigenic variation.
Despite the N-terminal domain structure of one VSG, MITat1.2, having been solved and published in 1990, the structure of a complete VSG has proven difficult to obtain. In fact, at the start of this study, structural data was only available for the individual domains of two of the large family of VSGs, MITat1.2 and ILTat1.24.
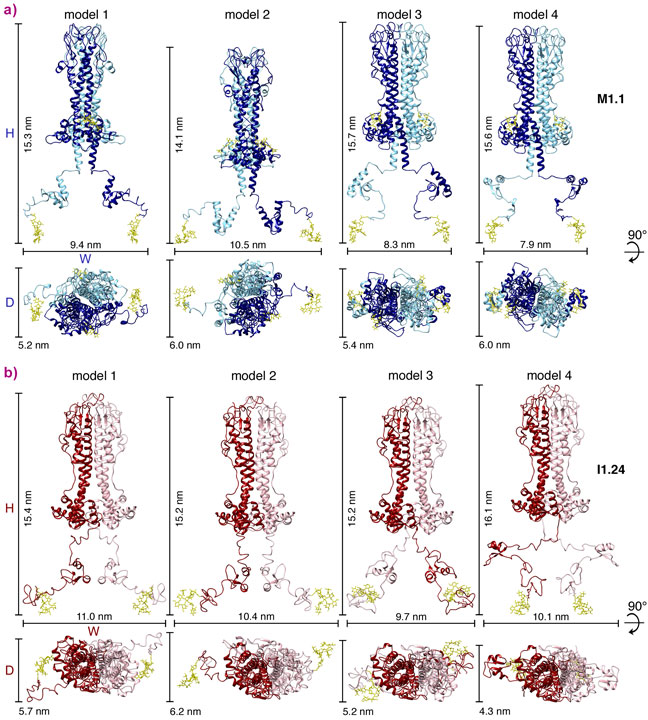
Here, the domain structures of a third VSG, MITat1.1, were solved by X-ray crystallography (N-terminal domain) and NMR spectroscopy (C-terminal domain). Subsequently, SAXS experiments on purified full-length VSG were conducted at beamline BM29. In addition, SAXS data for VSG ILTat1.24 were acquired, for which high-resolution N- and C-terminal domain structures were already available. The SAXS experiments allowed low-resolution molecular envelopes to be generated of both VSGs into which the high-resolution domain structures could be fitted. By doing this, structural models for both VSGs were obtained that fell into one of two groups (Figure 1), suggesting the existence of a compact more elongated form and a relaxed form that covers a larger area of the cell surface. Single molecule tracking measurements in supported lipid bilayers revealed the existence of two distinct, freely diffusing VSG populations, thus supporting the existence of two main conformations of VSGs on the cell surface.
The combination of SAXS data and single molecule diffusion experiments suggests that the adoption of two main VSG conformations allows surface shielding even at varying protein densities. Furthermore, high mobility of VSGs could be supported, even in the presence of obstacles and confinement within the plane of the plasma membrane.
Principal publication and authors
Structural basis for the shielding function of the dynamic trypanosome variant surface glycoprotein coat, T. Bartossek (a), N.G. Jones (a), C. Schäfer (b), M. Cvitković (c,d), M. Glogger (a), H.R. Mott (e), J. Kuper (b), M. Brennich (f,g), M. Carrington (e), A.-S. Smith (c,d), S. Fenz (a), C. Kisker (b), M. Engstler (a), Nature Microbiology 2, 1523–1532 (2017); doi: 10.1038/s41564-017-0013-6.
(a) Department of Cell and Developmental Biology, University of Würzburg (Germany)
(b) Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg (Germany)
(c) Institut für Theoretische Physik and the Excellence Cluster: Engineering of Advance Materials, Friedrich-Alexander-Universität Erlangen-Nürnberg, Erlangen (Germany)
(d) Group for Computational Life Sciences, Division of Physical Chemistry, Ruđer Bošković Institute, Zagreb (Croatia)
(e) Department of Biochemistry, University of Cambridge (UK)
(f) ESRF
(g) European Molecular Biology Laboratory, Grenoble (France)