- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2012
- Structural biology
- Structural analysis reveals features of the spindle checkpoint kinase Bub1-kinetochore subunit Knl1 interaction
Structural analysis reveals features of the spindle checkpoint kinase Bub1-kinetochore subunit Knl1 interaction
The spindle assembly checkpoint, or mitotic checkpoint, coordinates mitotic timing with chromosome-spindle interactions during mitosis, restricting mitotic exit to cells that have achieved bi-orientation of all their chromosomes [1]. Cells in which the checkpoint is artificially inactivated undergo precocious mitotic exit in the presence of unattached or incorrectly attached chromosomes. Alterations of checkpoint function might be relevant for tumour development, possibly by rendering cells more susceptible to the development of aneuploidies and to consequent genetic instability.
The Bub1 (budding uninhibited by benzimidazole 1) serine/threonine kinase is a conserved component of the spindle assembly checkpoint. During mitosis, Bub1 localises to kinetochores, large scaffolds that bridge chromosomes to spindle microtubules to ensure faithful chromosome segregation. It is believed that kinetochore recruitment of Bub1, where the protein displays slow exchange dynamics that probably reflect its specific activities there, is essential for its function. In our studies, we decided therefore to investigate the mechanism of kinetochore recruitment of the Bub1 kinase.
Kinetochore recruitment of Bub1 has been proposed to rely on the interaction of the Bub1 tetratricopeptide repeats (TPR) with a short sequence motif of the outer kinetochore protein Knl1 known as KI1 [2,3]. To test this possibility formally, we reconstituted a construct of the TPR repeats of Bub1 with the KI1 motif of Knl1 and crystallised it. X-ray diffraction data from a single crystal were collected at beamline ID14-1 to a resolution of 2.6 Å in spacegroup P21. Four independent monomers were placed in the asymmetric unit by molecular replacement using the model of the Bub1 TPRs from S. cerevisiae (PDB ID 3ESL) as a search probe [4].
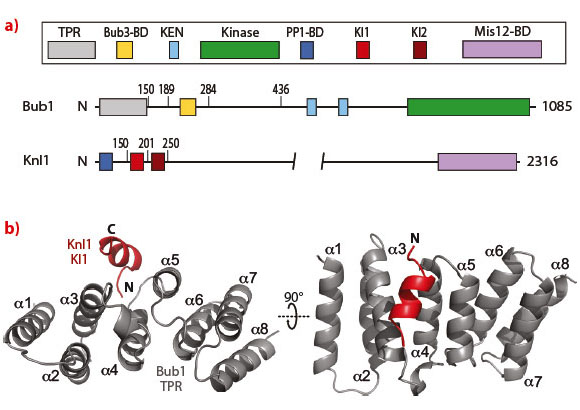
The crystal structure revealed that the interaction of Bub1 with Knl1 develops along the convex surface of the TPR assembly (Figure 19). Alanine mutations at residues Phe75, Asn79, Gln84 and Phe85 of Bub1 led to complete disruption of the Bub1:KI1 interaction in vitro. We generated stable inducible cells expressing EGFP-tagged wild type Bub1 and a mutant carrying the alanine mutations. Surprisingly, kinetochore recruitment of the Bub1(4A) mutant appeared to be largely normal. This observation confuted the hypothesis that the interaction of the TPR region of Bub1 with the KI1 motif of Knl1 is important for kinetochore recruitment of Bub1.
 |
|
Fig. 19: a) Schematic representation of the domain structures of Bub1 and Knl1. b) Cartoon diagram of the TPR (grey):KI1 (red) interaction. The KI1 peptide binds the convex surface of the TPR helical array. |
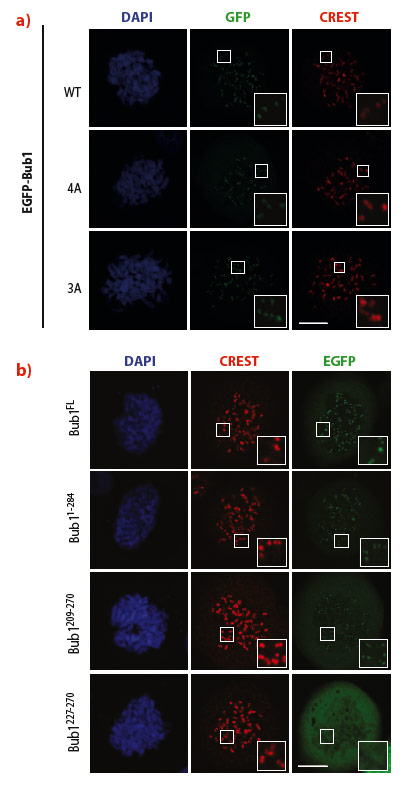
Next, we sought to identify the minimal segment of Bub1 required for kinetochore recruitment. After showing that residues 190 and 284 of human Bub1 are sufficient for kinetochore recruitment, we refined our analysis by expressing additional constructs, including Bub1(209-270) and Bub1(227-270). The latter construct matched almost exactly the segment of budding yeast Bub1 that was previously co-crystallised with Bub3 [4], while the former is preceded by an 18-residue N-terminal extension of unknown function. While Bub1(209-270) localised robustly to kinetochores, Bub1(227-270) was unable to reach kinetochores (Figure 20). These results suggest that efficient kinetochore recruitment requires the Bub3 binding domain and a short N-terminal extension.
 |
|
Fig. 20: a) Two combinations of multiple alanine point mutants of Bub1, indicated by 4A and 3A, leave kinetochore recruitment of Bub1 unaltered. CREST is a kinetochore marker. The inset shows enlarged kinetochores. b) Identification of a minimal kinetochore-binding segment of Bub1. Bub1(209-270) localised normally to kinetochores, whereas Bub1(227-270) failed to be recruited. |
Our studies exploited the crystal structure of the Bub1-KI1 complex to design a set of simple and insightful functional experiments that led us to question a previously proposed mechanism for kinetochore recruitment of Bub1. On the basis of these negative results, we went on to identify the precise determinants for Bub1 kinetochore recruitment [4].
Principal publication and authors
V. Krenn (a), A. Wehenkel (a,b), X. Li (a), S. Santaguida (a) and A. Musacchio (a,b), J Cell Biol. 196, 451-467 (2012).
(a) Department of Experimental Oncology, European Institute of Oncology, Milan (Italy)
(b) Department of Mechanistic Cell Biology, Max Planck Institute of Molecular Physiology, Dortmund (Germany)
References
[1] A. Musacchio and E.D. Salmon, Nat Rev Mol Cell Biol 8, 379–393 (2007).
[2] T. Kiyomitsu, C. Obuse and M. Yanagida, Dev Cell 13, 663–676 (2007).
[3] T. Kiyomitsu, H. Murakami & M. Yanagida, Mol Cell Biol 31, 998–1011 (2011).
[4] N.A. Larsen, J. Al-Bassam, R.R. Wei and S.C. Harrison, Proc Natl Acad Sci USA 104, 1201–1206 (2007).



