- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2010
- Soft condensed matter
- New results may help stop age-related macular degeneration at the molecular level
New results may help stop age-related macular degeneration at the molecular level
Age-related macular degeneration, or AMD, is painless but affects the macula, the part of the retina that allows one to see fine detail. The earliest hallmark of AMD is the appearance of protein, lipid and zinc deposits under the retinal pigment epithelial cells. These yellowish deposits are usually discovered by an ophthalmologist during routine eye tests and are commonly known as “drusen”. AMD is the number one cause of vision loss in the elderly, causing 57% of blindness in the UK alone.
A key insight into the molecular beginnings of AMD was obtained by determining how two key proteins interact normally to prevent the onset of the condition. By the use of a multidisciplinary strategy that included X-ray scattering at beamline ID02, researchers at UCL clarified for the first time how an abundant blood protein called complement Factor H (of innate immunity) binds with another protein called C-reactive protein, to reduce the potentially damaging effects of an overactive immune system. In the eye, during ageing processes, cells will die naturally for all sorts of reasons. In conditions of high inflammation, at the same time as the levels of C-reactive protein in the retina increase dramatically, uncontrolled complement activity may cause damage to the retina. It is mostly the debris of broken up retinal cells that is deposited as drusen. The role of C-reactive protein is to coat these damaged cells, while the importance of factor H is to slow down high inflammation.
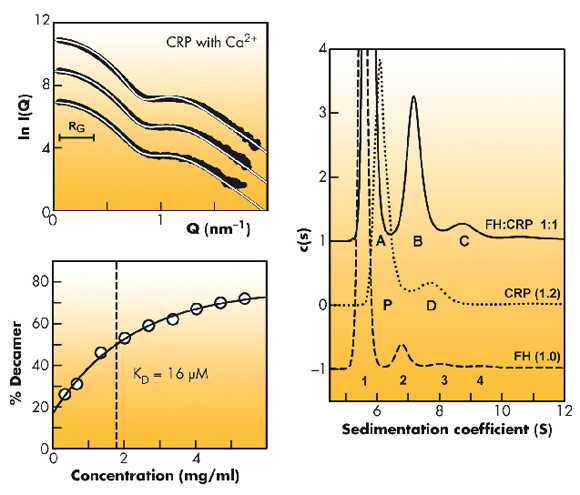
The interaction between factor H and C-reactive protein was previously hindered by a lack of understanding of their solution properties. For unbound C-reactive protein, X-ray scattering, analytical ultracentrifugation and surface plasmon resonance showed that pentamers and decamers were formed in solution with the same dissociation constant at 16-23 µM (Figure 72). The study emphasised the importance of working with 2 mM calcium and 140 mM NaCl in order to avoid protein denaturation, especially after the immobilisation procedure used in surface plasmon resonance. For Factor H, scattering and ultracentrifugation has already shown that Factor H formed dimers, trimers and higher oligomers in addition to the major monomer form.
On mixing the two proteins together, ultracentrifugation showed that decamer and dimer/trimer formation in the two proteins was suppressed (Figure 72), proving that the proteins interact, but only in the right buffer with undenatured proteins. Surface plasmon resonance confirmed the interaction with a dissociation constant of 4 µM. Scattering showed conformational changes that occurred when the proteins were mixed. Two molecules of C-reactive protein bind to Factor H at two different sites. It was then found that a common genetically modified form of Factor H (with His402) does not bind to C-reactive protein as well as the normal form (with Tyr402) (Figure 73). People with modified Factor H are more vulnerable to an immune system attack in the eye and, thus, drusen may build up.
 |
|
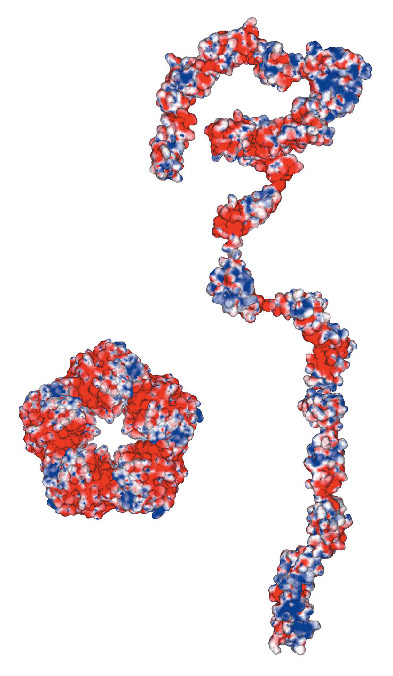
Fig. 73: Electrostatic surfaces of complement factor H with 20 SCR domains, a regulator of the innate immune system, and the C-reactive protein pentamer, the classic acute phase protein. Red denotes acidic regions while blue denotes basic regions. Both proteins are drawn to the same scale. The factor H solution structure was determined by constrained X-ray scattering modelling, while the C-reactive protein structure was determined from protein crystallography. |
While there is no known cure for AMD, existing therapies aim to treat the symptoms and delay progression. The new biophysical results revealed from this work clarified one of the major molecular interactions that had proven hard to unravel by other immune techniques.
Principal publications and authors
A.I. Okemefuna, L. Stach, S. Rana, A.J. Ziai Buetas, J. Gor, R. Nan, A. Miller and S.J. Perkins, The Journal of Biological Chemistry 285, 1041-1052 (2010); The Journal of Biological Chemistry 285, 1053-1065 (2010).
Research Department of Structural and Molecular Biology, University College London (UK)