- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2010
- Structural biology
- The structure of the nuclease domain of human cytomegalovirus DNA packaging terminase
The structure of the nuclease domain of human cytomegalovirus DNA packaging terminase
Human cytomegalovirus (HCMV), one of the eight human herpesviruses, has a high prevalence around the world. In healthy hosts, infection is usually silent but it has life-threatening consequences in immunocompromised individuals. The main groups at risk of HCMV infection are newborn and young children, organ-transplant receivers, patients treated for cancer and people co-infected with HIV. In these risk groups, symptoms can entail loss of hearing and vision, mental retardation and cerebral palsy.
HCMV replicates its DNA via concatamers, a long molecule of DNA containing several copies of the genome. This DNA has to be cut into single genomes and packaged inside the viral capsid. This packaging process is performed by the terminase complex, which in HCMV includes at least, two proteins UL89 and UL56. The observation that DNA packaging processes do not occur in mammalian cells implies that the viral proteins involved might be specific targets for antiviral therapy. However, despite interest in herpes virus terminases as drug targets, no structural information is available for these proteins. Here we focussed on the structural and functional analysis of one component of the HCMV terminase complex, the UL89 protein.
First, we attempted, without success, to express the full-length UL89 in several hosts. In all the cases, the level of expression was very low and the protein expressed was insoluble. Next, to find a soluble fragment of UL89, we predicted domain boundaries using a number of bioinformatic tools; however, all the fragments found were again insoluble. At this point, we attempted a high throughput expression method, ESPRIT, which entails the search for expression of soluble protein fragments by random incremental truncation [1]. Using this approach to test 18432 different gene constructs, we found a soluble domain of UL89, corresponding to the last 257 amino acids of the protein, which we termed UL89-C.
 |
|
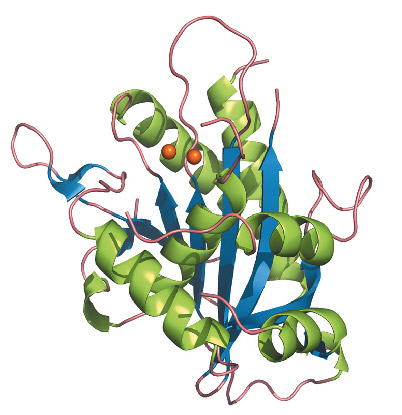
Fig. 119: Overall structure of UL89-C in a ribbon representation. |
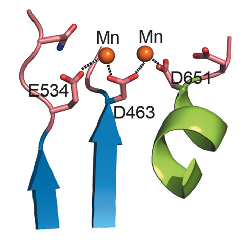
UL89-C was subsequently purified and crystallised, and its crystal structure solved using X-ray diffraction data collected at beamlines ID14-2 and BM16. The structure of UL89-C is composed of a central eight-stranded β-sheet surrounded by six α-helices (Figure 119). Although there are some extra decorations, the fold of UL89-C corresponds to that of RNAse H, an enzyme that removes RNA primers from Okazaki fragments. Other proteins with this fold are E. coli Ruv-C resolvase, HIV and ASV integrases, some transposases and other nucleases [2]. All these proteins share a basic fold of a five-stranded β-sheet surrounded by α-helices on both sides. The order and orientation of the strands within the β-sheet is conserved: 3, 2, 1, 4 and 5, one of them being anti-parallel to the other four. An anomalous difference Fourier map, calculated with diffraction data collected at the Mn-K absorption edge at beamline ID29, confirmed that the active site of UL89-C contains two Mn2+ ions approximately 4 Å apart. The first Mn2+ is coordinated by Asp463 and Glu534 and the second by Asp463 and Asp 651 (Figure 120). These residues are fully conserved in all human herpes virus terminases.
 |
|
Fig. 120: Detailed view of the active site area of UL89-C. |
To demonstrate that UL89-C encodes the nuclease activity of the terminase, we performed a nuclease assay using plasmidic DNA. We tested the reaction in the presence of various divalent ions. Only with Mn2+ was total degradation of DNA observed. This finding indicates that nuclease activity is strongly dependent on this cation. We also checked the nuclease activity with mutants of the residues that coordinate the Mn2+ ions and confirmed that they are essential for the function of the protein.
HIV integrase, which is a validated anti-HIV drug target, shares its basic fold with UL89-C and has a similar active site. Prompted by this structural similarity, we tested the inhibitory properties towards UL89-C of a number of known HIV integrase inhibitors. Raltegravir [3], an anti-HIV drug approved by the FDA in October 2007 and widely used in antiretroviral therapy for AIDS patients, was active against the UL89-C terminase domain at low micromolar levels. This observation opens the way for the development of antiviral molecules against herpesviruses using the nuclease domain of the terminase as a target.
Principal publication and authors
M. Nadal (a,b), P.J. Mas (c), A.G. Blanco (a,b), C. Arnan (a,b), M. Solà (b), D.J. Hart (c) and M. Coll (a,b), Proc Natl Acad Sci USA 107, 16078-16083 (2010).
(a) Institute for Research in Biomedicine (IRB Barcelona) (Spain)
(b) Institut de Biologia Molecular de Barcelona (CSIC) (Spain)
(c) EMBL Grenoble (France)
References
[1] H. Yumerefendi, F. Tarendeau, P.J. Mas and D.J. Hart, J Struct Biol 172, 66-74 (2010).
[2] W. Yang and T.A. Steitz, Structure 3, 131-134 (1995).
[3] V. Summa et al., J. Med. Chem 51, 5843-5855 (2008).



