- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2010
- Structural biology
- An original Cu(I)-coordination shell in a small bacterial metalloprotein identified by XAS and NMR spectroscopy
An original Cu(I)-coordination shell in a small bacterial metalloprotein identified by XAS and NMR spectroscopy
Living organisms are continuously exposed to environmental stress from sources such as toxic compounds, radiation, and high temperatures. During evolution, they have adapted to the specific environment in which they live and have learned to protect themselves against sources of stress by developing resistance mechanisms. One well-known class of toxic products is heavy metals that, although naturally occurring in the environment, may be released in huge quantities through anthropogenic processes.
Cupriavidus metallidurans CH34 is a bacterium that is resistant to high concentrations of many heavy metals and is a model for studying metal resistance mechanisms in microorganisms. Copper resistance of C. metallidurans CH34 involves more than 19 proteins. One of these is the metallochaperone CopK, a small periplasmic protein that is found in significant amounts under copper stress. We have previously determined the solution structure of apo-CopK [1] which has been shown to bind Cu(I) and Cu(II) in a cooperative way [2]. Recently, we determined the solution structure of the Cu(I)-bound CopK protein and characterised its Cu(I) binding site. A thus far unique Cu(I) ligand sphere, formed by four thioether groups belonging to four methionines from the same protein chain, was discovered.
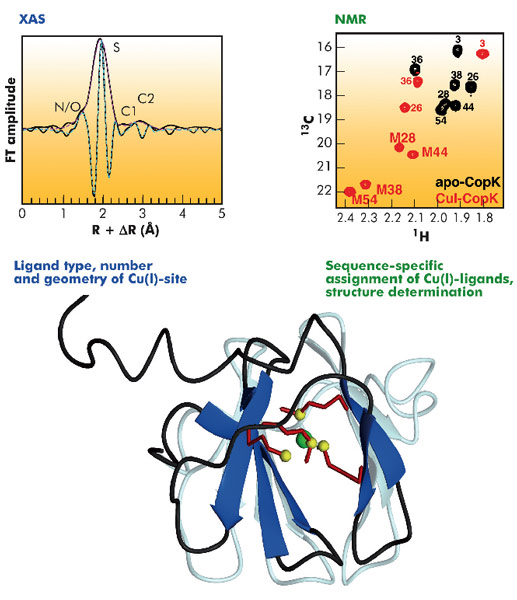
The three-dimensional structure of CopK was determined from NMR data (Figure 118). However, the Cu(I) ion is NMR-silent and no geometric information on the metal site could be obtained. A cryo-cooled protein solution containing Cu(I)-CopK was therefore studied by XAS, and spectra around the Cu K-absorption edge collected on the BM30B (FAME) beamline. The resulting XANES and EXAFS spectra were in accordance with a coordination sphere formed by four sulphur atoms and the fit of the EXAFS data suggested a Cu(I)-S distance of 2.31 Å, a value typical for tetragonal sites. This result was surprising as in most small periplasmic bacterial Cu(I)-binding metallochaperones, Cu(I) is bound in trigonal sites by a combination of sulphur and nitrogen ligands. In addition, a tetrathioether Cu(I) site has so far only been observed in a single protein in which the Cu(I) ion belongs to a copper-molybdenum (MoS2CuS2Mo) cluster. Our findings were further corroborated by the observation of chemical shift differences in 1H-13C correlation NMR spectra occurring upon Cu(I)-binding. Four peaks, corresponding to methionine methyl groups were significantly affected by the addition of Cu(I) to the protein. This observation suggests that the chemical shift of these carbons allows the detection of methionine-bound Cu(I) and thus the sequence-specific identification of the Cu(I) ligands.
The structure of Cu(I)-CopK bridges the gap between what was previously known on the Cu(I)/Cu(II)-binding cooperativity and the structures of CopK previously reported. Accommodation of Cu(I) in the tetrathioether site is associated with an important conformational change of the C-terminal region of CopK. We propose that this reorientation is also required for the formation of the Cu(II)-specific site, a hypothesis consistent with the much higher Cu(II) affinity of Cu(I)-CopK compared to apo-CopK.
Principal publication and authors
G. Sarret (a), A. Favier (b), J. Covès (b), J.-L. Hazemann (c), M. Mergeay (d) and B. Bersch (b), J. Am. Chem. Soc. 132, 3770–3777 (2010).
(a) LGIT, Grenoble (France).
(b) Institut de Biologie Structurale, Grenoble (France).
(c) Institut Néel, Grenoble (France).
(d) SCK-CEN, Mol (Belgium)
References
[1] B. Bersch et al., J. Mol. Biol. 380, 386-403 (2008).
[2] L.X. Chong et al., J. Am. Chem. Soc. 131, 3549-3564 (2009).