- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2009
- Soft condensed matter
- Crystal structure of A-amylose by synchrotron microdiffraction
Crystal structure of A-amylose by synchrotron microdiffraction
The molecular and crystal structure determination of low molecular weight compounds is carried out with great precision on a routine basis. However, for polymer crystals, resolution of the structure is still a major challenge since it requires a number of intuitive guesses together with trial-and-error methods [1]. Indeed, many small molecules are susceptible to crystallisation as millimetre or sub-millimetre-sized crystals, yielding extensive diffraction data sets, with frequently more than 10 to 20 observed reflections per independent atom, the minimum ratio that is required for a proper structure refinement. This is not the case for most polymers for which only crystals with dimensions ranging from sub-micrometre to the micrometre range can be obtained. These crystals are far too small to yield substantial X-ray datasets when conventional X-ray diffractometers are used. In most cases, one relies on the study of crystalline fibres by X-ray diffraction and single-crystals by electron diffraction techniques. Both of the methods have severe challenges when compared with single-crystal X-ray diffraction: unambiguous deconvolution of fibre diffraction diagrams into three dimensional data; and the difficult exploitation of the electron diffraction intensities for crystal structure determination since one needs to subtract dynamic contributions from electron diffraction intensities before any further analysis.
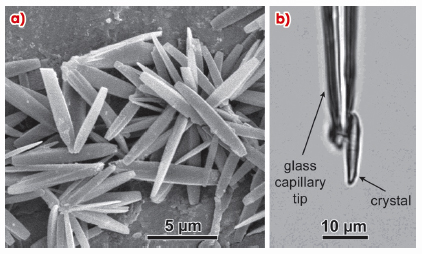
The recent development of microfocus beamlines at third generation synchrotron radiation sources, such as the ESRF [2], has greatly improved the situation as novel X-ray optical systems allow the production of focal spots with diameters in the micrometre and sub-micrometre sizes. In addition, goniometers for keeping a single crystal in the centre of micrometre-sized beams have also been developed. Thus, when applied to very small crystalline objects such as a polymer single crystal, one can envisage recording X-ray datasets that are more complete than those recorded in electron diffraction experiments, and therefore susceptible to yield improved structural models. Here we show how a full three-dimensional dataset was collected on microcrystals of A-amylose (Figure 77) and used to refine the structure of this amylose allomorph. To our knowledge, this is the first polymer structure determined using this technique on such small polymer single crystals. Getting an accurate atomic description of the A-amylose crystal is important since these crystals constitute part of the cereal starches, a food product of capital importance.
 |
|
Fig. 77: a) SEM image of A-amylose crystals; b) One of the crystals used for data collection. |
Experiments were performed at beamline ID13. The crystal structure was solved using a molecule replacement technique. The final refinement, which was done in the isotropic approach of thermal parameters, gave R1 and R1free values of 0.17 and 0.22 respectively.
 |
|
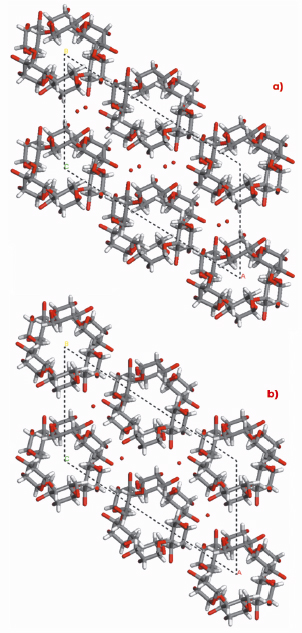
Fig. 78: Structures of A-amylose: a) present and b) past. |
In agreement with the earlier structure based on X-ray fibre powder and electron diffraction data, our A-amylose structure also consists of intertwined 6-fold left-handed parallel double helices, packed in a parallel fashion and tightly nested into one another. Furthermore, the existence of two water molecules per maltotriosyl unit was revealed without ambiguity in the crystal structure of A-amylose, whereas, from density consideration, only one was proposed in the earlier structure. This new feature brings substantial difference, not only for the system of hydrogen bonds, which has to account for new possibilities, but also for a distortion of the helices and of their internal cavity. This feature is well illustrated in Figure 78.
References
[1] T. Tadokoro, in Structure of Crystalline Polymers; John Wiley and Sons: New-York, p 6 (1979).
[2] C. Riekel, J. Rep. Prog. Phys. 63, 233-262 (2000).
Principal publication and authors
D. Popov (a), A. Buléon (b), M. Burghammer (a), H. Chanzy (c), N. Montesanti (c), J-L. Putaux (c), G. Potocki-Véronèse (d) and C. Riekel (a), Macromolecules 42, 1167-1174 (2009).
(a) ESRF
(b) INRA, Nantes (France)
(c) Centre de Recherches sur les Macromolécules Végétales (CERMAV-CNRS), Grenoble (France)
(d) Laboratoire d’Ingénierie des Systèmes Biologiques et des Procédés (LISBP), INSACNRS, Toulouse (France)



