- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2009
- Soft condensed matter
- Biological imaging by coherent hard X-ray diffraction microscopy
Biological imaging by coherent hard X-ray diffraction microscopy
Imaging of biological material in three dimensions, such as an entire cell, deepens our insight into cellular biology. The highest possible resolution of the images is required as well as an experimental environment preserving the samples in their natural (hydrated) state without introducing structural artefacts, such as staining or sectioning, and radiation damage. Meeting these requirements is obviously a formidable task.
 |
|
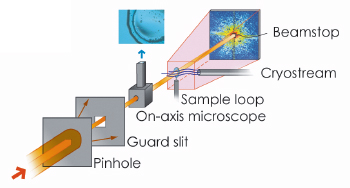
Fig. 69: Sketch of the experimental setup for bio-XDM at ID10C. |
X-ray diffraction microscopy (XDM also known as CXDI, coherent X-ray diffraction imaging) is an image reconstruction method where the coherent X-ray diffraction pattern of a sample is inverted to yield the real space structure [1]. This technique has potential to reach the goal of 3D biological imaging. Figure 69 shows the basic XDM experimental setup we employed in this study. The image reconstruction, r(r), is performed by recovering the phase of the diffracted intensity pattern, I(Q), through a phase retrieval algorithm [2]. Thanks to the high penetration power of X-rays, a cell can be imaged as a whole, without sectioning. Unlike most other imaging techniques the ultimate resolution is not determined by a lens (therefore XDM is often denoted as lensless imaging), but rather by the largest momentum transfer Q to which I(Q) can be precisely measured. For biological samples, radiation damage thresholds set the resolution limits. Nevertheless, soft X-ray experiments on dry unstained yeast cells have proven the feasibility of the method [3]. Hard X-rays are preferred in XDM since the convergence of the phasing algorithm, especially for three-dimensional reconstructions, is improved within the Born-approximation. However, it was unknown until recently whether the signal-to-noise ratio of scattered hard X-rays from unstained biological material in a hydrated state was sufficient for XDM.
 |
|
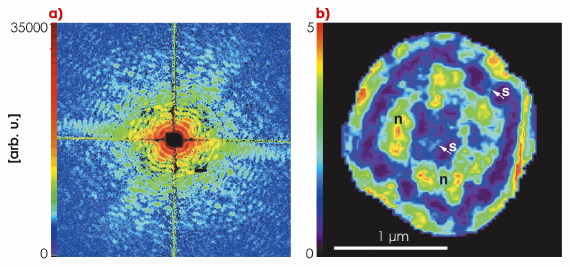
Fig. 70: a) Coherent diffraction pattern (speckle) of a hydrated D. radiodurans bacterial cell. b) The reconstructed image where sub-cellular nucleoid structures are clearly visible. |
Figure 70 shows the far-field diffraction patterns of unstained frozen-hydrated Deinococcus radiodurans bacteria measured at beamline ID10C (Troika) using 8 keV X-rays. Samples were rapidly plunge-frozen in the hydrated state to avoid structural artefacts from dehydration and to prevent radiation damage. Mounted on the diffractometer, ice contamination was avoided or reduced to a minimum by keeping the samples in a cold nitrogen gas environment provided by a cryogenic gas stream. Neither staining, nor sectioning was applied to the few-micrometre-thick bacterial cells. Thus the contrast of the diffraction pattern arises only from the cellular material. The spatial frequencies at the edge of the diffraction pattern correspond to a half-period of 25 nm resolution in real space. By filtering and phasing the diffraction pattern, it could be inverted as shown in Figure 70. Since this image is just one 2d projection of the three-dimensional cell structure, its interpretation may be difficult. Hence, an extension to 3d cryogenic bio-XDM is desirable, and this is now in progress.
References
[1] J. Miao, P. Charalambous, J. Kirz and D. Sayre, Nature 400, 342 (1999).
[2] J.R. Fienup, Appl. Opt. 21, 2758 (1982).
[3] D. Shapiro et al., PNAS 102, 15343 (2005
Principal publication and authors
E. Lima (a, b), L. Wiegart (a, b), P. Pernot (a), M. Howells (a, c), J. Timmins (a), F. Zontone (a), and A. Madsen (a), Phys. Rev. Lett. 103, 198102 (2009).
(a) ESRF
(b) NSLS-II, Brookhaven National Laboratory (USA)
(c) LBNL, Berkeley (USA)



