- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2003
- Macromolecular Crystallography
- Crystal Structure of the Retinoblastoma Tumour-suppressor Protein bound
Crystal Structure of the Retinoblastoma Tumour-suppressor Protein bound
The first tumour-suppressor protein to be identified was the product of the retinoblastoma gene (pRb). Loss of its function contributes to the development of a majority of human malignancies [1]. pRb plays important roles in regulating the cell-cycle, apoptosis and differentiation, and all of these activities are pertinent to its role as a tumour suppressor. The growth-inhibitory effects of pRb are dependent on its regulation of the E2F family of transcription factors. In order to better understand the regulation of the E2F transcription factor by pRb, we have determined the crystal structure of the pRb pocket domain (pRbAB) bound to residues 409-426 of E2F-1. The AB-pocket of pRb is the major focus of tumourigenic mutations in the protein and the E2F(409-426) fragment represents the core of the pRb-binding region of the transcription factor.
 |
|
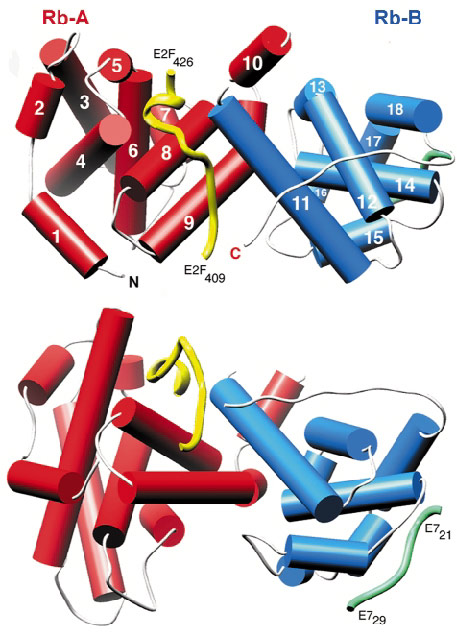
Fig. 25: The structure of RbAB/E2F(409-426), shown in two orthogonal views (drawn with Ribbons). The helices of the A domain are shown as red cylinders and those of the B domain as blue cylinders. The main-chain trace of E2F and E7 (from PDB entry 1GUX) are shown as yellow and green worms respectively. |
Initial attempts at data collection, using synchrotron X-ray sources, were hindered by very high crystal mosaicity and poor data scaling and reduction. Finally, a good quality dataset was collected to 2.6 Å resolution using the microfocus diffractometer on beamline ID13. At the time this was the only such device installed at a synchrotron source. The structure was solved by molecular replacement and the E2F peptide could be readily located in the initial electron density maps. The structure shows that the packing of the A and B domains generates a waist-like interface groove into which E2F(409-426) binds in a largely extended manner (Figure 25). A high proportion of the hydrogen bond interactions between the two molecules involve the side chains of conserved pRb residues interacting with the main chain of E2F.
Some oncogenic viruses, such as human papillomavirus (HPV) code for proteins that disrupt pRb/E2F interaction. In the case of HPV this function is mediated by the small, zinc-binding protein E7 that binds to pRb via the characteristic LxCxE motif of the oncoprotein. The crystal structure reveals that the core binding site of E2F is located more than 30 Å away from the E7(21-29) binding site (green worm in Figure 25). To examine the mechanism of how E7 targets the pRb/E2F complex, a series of biochemical experiments have been carried out using intact HPV E7 and a construct of E2F that contains both the marked box and the core binding region. The data demonstrate that intact E7 protein binds at least 15-fold tighter to pRb than a short LxCxE based E7 peptide. These results also reveal that the tight biding of the HPV oncoproteins to pRb prevents subsequent interactions with the marked box region of E2F but does not affect binding of the tumour suppressor to the core binding-region of E2F.
With the detailed molecular description of the interactions between E2F and pRB, it should be possible to design site-specific mutants of pRb that no longer bind E2F, but whose other properties are unaltered. Such mutated pRb would be a valuable tool for cell biology experiments aimed at probing the function of pRb/E2F interactions. There are some tumours, such as pancreatic carcinoma, where over-expression of functional pRb appears to be detrimental to clinical treatment [2]. Given our description of the molecular interactions between E2F and the A/B interface of pRb we can develop compounds that bind to pRb and inhibit complex formation. Such a compound, administered in parallel with conventional chemotherapy, may offer a means of treatment for pancreatic cancer and perhaps related diseases.
References
[1] R.A. Weinberg, Cell 81, 323-30 (1995).
[2] T. Plath, M. Peters, K. Detjen, M. Welzel, Z. von Marschall, C. Radke, B. Wiedenmann, and S. Rosewicz, J Natl. Cancer Inst. 94, 129-142 (2002).
Principal Publication and Authors
B. Xiao (a), J. Spencer (a), A. Clements (b), N. Ali-Khan (a), S. Mittnacht (c), C. Broceño (c), M. Burghammer (d), A. Perrakis (e), R. Marmorstein (b) and S.J. Gamblin (a), Proc. Natl. Acad. Sci. USA 100, 2363-2368 (2003).
(a) National Institute for Medical Research (UK)
(b) University of Pennsylvania (USA)
(c) Institute of Cancer Research (UK)
(d) ESRF
(e) The Netherlands Cancer Institute (The Netherlands)



