- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2003
- Macromolecular Crystallography
- The Structure of Plant Photosystem I
The Structure of Plant Photosystem I
The conversion of solar energy into chemical bonds by the photosynthetic machinery of plants, green algae and cyanobacteria is essential for the survival of all higher life forms. The primary event of this process - light induced charge translocation - is catalysed by photosystem I (PSI) and photosystem II (PSII), two multisubunit protein complexes embedded in the thylakoid membrane that act in series. PSI harnesses light energy to transfer an electron from the soluble copper protein, plastocyanin, on one side of the membrane to ferredoxin and eventually NADPH on the opposite side. In the dark, NADPH (and ATP) provides the energy for assimilation of CO2 into carbohydrates. PSI is composed of a 12 subunit reaction center and a 4 subunit membranal light-harvesting antenna (LHCI) that captures light and channels its energy to the reaction center. LHCI is exclusively found in plants and green algae and consists of four different but homologous chlorophyll a/b binding proteins, termed Lhca1-4.
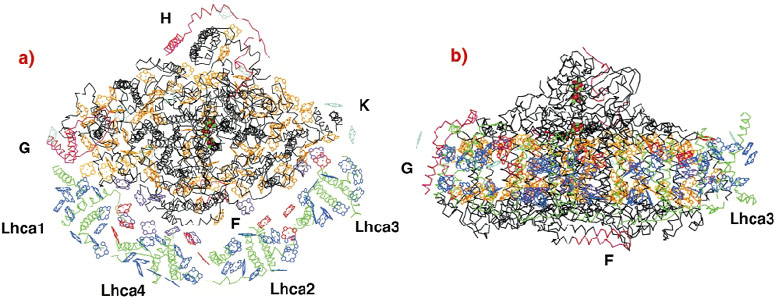
We have determined the crystal structure of the entire PSI complex from a higher plant (pea) to 4.4 Å resolution [1]. Phases were obtained by the MIRAS (multiple isomorphous replacement including anomalous scattering) method using several heavy-atom derivatives and the anomalous signal of intrinsic Fe-S clusters at 0.93 Å wavelength. With a mass of 530 KDa, 45 transmembrane helices and 172 co-factors (mostly chlorophylls), this is the largest and most intricate membrane assembly solved by X-ray crystallography and the first one from higher plants (Figure 22). The high intensity of beamline ID14-4 proved crucial for obtaining the reported resolution. All data used for this structure solution were collected there.
The four LHCI proteins assemble into two dimers, forming a half-moon shaped belt that docks to the subunit F side of the reaction center. The association of the reaction center with the LHCI belt is both weak and asymmetric (Figure 22a). Only one of the four Lhca proteins firmly attaches to the reaction center through interactions involving transmembrane helices and may serve as an "anchor" for the assembly of the other monomers. Within the membrane, the Ca backbone of LHCI maintains a distance of 20 Å from most parts of the reaction center. Thus, LHCI chlorophylls are positioned 18 Å from the nearest reaction center chlorophyll except at three regions where much shorter inter-pigment distances are observed and which may play an important role in energy migration. The arrangement of the ~90 chlorophylls within the reaction center in plants is similar to that in the cyanobacterial counterpart [2]. Apparently, adaptation of the reaction center chlorophyll arrangement to receiving energy from LHCI required the addition of only an extra 10 chlorophylls at strategic positions between LHCI and the reaction center.
Formation of the LHCI dimers and the interaction between the dimers does not involve transmembrane helices. The solvent exposed C and N termini of the light-harvesting protein monomer attach to solvent-exposed loops of its neighbouring monomer (Figure 22). "Linker chlorophylls" between LHCI monomers facilitate energy migration along the LHCI belt. This mode of binding maximises the number of LHCI chlorophylls facing the reaction center. Furthermore, we suggest that this dimerisation mode, together with the above-described interactions between LHCI and the reaction center serve the need of PSI to adjust LHCI composition according to varying environmental conditions.
The structure reveals further details concerning, for example, interactions of PSI with other light-harvesting proteins (LHCII) and with plastocyanin. It provides a framework for investigating both the mechanisms regulating the activity and composition of PSI's numerous subunits and the evolutionary forces that shaped the photosynthetic apparatus of terrestrial plants.
References
[1] A. Ben-Shem, N. Nelson, and F. Frolow, Acta Crystallographica Section D-Biological Crystallography 59, 1824-1827 (2003).
[2] P. Jordan, P. Fromme, H.T. Witt, O. Klukas, W. Saenger, and N. Krauss, Nature 411, 909-917 (2001).
Principal Publication and Author
A. Ben-Shem, F. Frolow, and N. Nelson, Nature, 426, 630-635 (2003).
Tel Aviv University (Israel)