- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2001
- Life Sciences
- Antibiotics Targeting Ribosomes
Antibiotics Targeting Ribosomes
Analysis of high-resolution structures of complexes of antibiotics with ribosomal particles sheds light on antibiotic selectivity and illuminates various modes of action, from reduction of decoding accuracy via limiting conformational mobility, to interference with substrate binding and hindrance of the progression of growing proteins. Their interactions and the lack of major conformational rearrangements associated with antibiotic binding, support the suggestion that the ribosome provides a framework for peptide bond formation, rather than enzymatic activity.
Resistance to antibiotics is a significant problem in modern therapeutics. Ribosomes of pathogenic bacteria are major targets for antibiotics. Ribosomes are a cellular organelle catalysing the translation of genetic code into proteins. They are protein/RNA assemblies arranged in two subunits that associate for performing protein biosynthesis. The large subunit (1.5 megaDa, 3000 nucleotides in two RNA chains and ~35 proteins) creates the peptide bonds and provides the path for emerging nascent proteins. The smaller subunit (0.85 megaDa, 1500 nucleotides in one RNA chain and ~20 proteins) has key roles in controlling the fidelity of codon-anti-codon base-pairing and in initiating the biosynthetic process.
The high-resolution structures of ribosomal subunits from two pathogen-models [1], obtained recently by bright synchrotron radiation, were used as a reference that allowed unambiguous localisation of several antibiotics. Among those reported here, six were clinically relevant and one was of no clinical use. All were found to bind primarily to ribosomal RNA and their binding did not cause major conformational changes.
Small subunit antibiotics: Tetracycline was found to be a multi-site antibiotic with inhibitory action that stems from its interference with A-site tRNA binding. Edeine, a universal agent, inhibits the initiation of protein synthesis by linking critical features for tRNA, IF3 and mRNA binding, thus imposing constraints on ribosomal mobility that accompany the translation process (Figure 10). Its universality implies conservation of structural elements important for initiation.
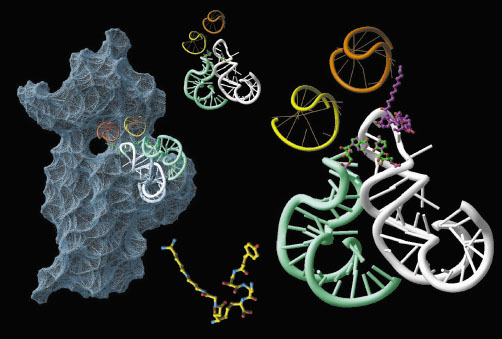 |
Fig. 10: (Left) The small ribosomal subunit. The mRNA path and the P-(orange) and E-(yellow) sites are shown. The RNA features that are "frozen" by edeine are highlighted in white and cyan. In the assembled ribosome the large subunit will face the left side of the particle. (Middle) top: the free edeine binding site. Bottom: the structure of edeine. (Right) Detailed view of edeine (purple) binding site. Note the newly formed base pair (green). |
Large subunit antibiotics: Chloramphenicol targets the peptidyl transferase cavity close to the amino acceptor group of tRNA. Clindamycin interferes with substrate binding and physically hinders the path of the growing peptide chain. The macrolides erythromycin, clarithromycin and roxithromycin bind to the entrance of the protein exit tunnel and block the progression of nascent proteins (Figure 11). Interestingly, none of these antibiotics binds to the nucleotides assigned to be crucial for the catalytic mechanism of the ribosome that was proposed based on the 2.4 Å structure of the Haloarcula marismortui large subunit [2].
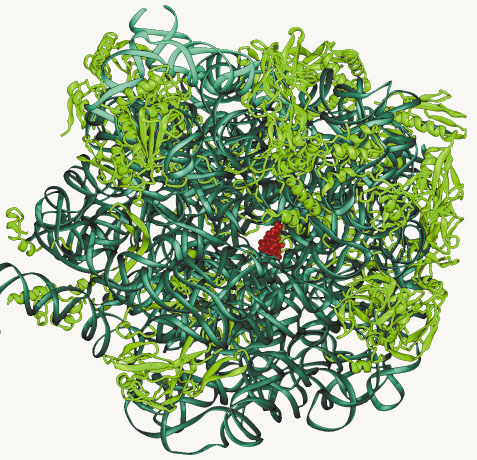 |
Fig. 11: The position of erythromycin (red) within the large ribosomal subunit - RNA (dark green), the proteins (light green). The view is from the active site into the protein exit tunnel. |
Comparative studies have helped to identify elements that may confer drug selectivity (e.g. Figure 12). The antibiotics modes of interactions and the preservation of the active-site conformation, favour the suggestion that the peptidyl transferase center serves as a template for proper positioning of tRNAs to allow for spontaneous, rather than enzymatic, creation of peptide bonds. The ribosomal components constructing the frame for accurate positioning of the tRNA molecules may include proteins, CTC, L27 and L16.
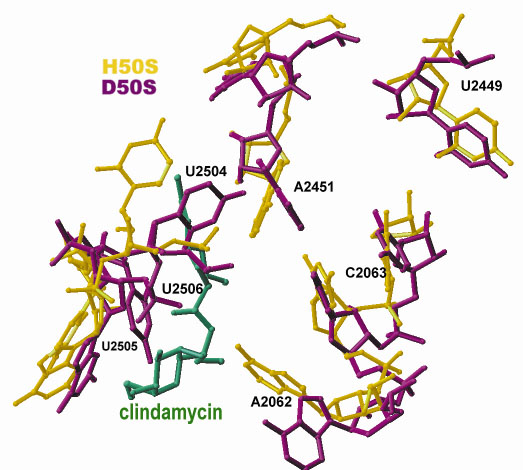 |
Fig. 12: Clindamycin binding site shown on a superposition of the backbone of the peptidyl transfer ring of a eubacterial pathogen model (D50S) and of its archeal counterpart (H50S) which serves as a model for eukaryotes (E. coli numbering scheme). |
Antibiotics targeting ribosomes are excellent tools for studying ribosomal function and for understanding mechanisms of drug action. Analysis of their modes of action should lead to structure-based design of improved antibiotics.
References
[1] F. Schluenzen, A. Tocilj, R. Zarivach, J. Harms, M. Gluehmann, et al., Cell, 102, 615-623 (2000).
[2] P. Nissen, J. Hansen, N. Ban, P.B. Moore and T.A. Steitz, Science 289, 920-930 (2000).
Principal Publications and Authors
F. Schluenzen (a), R. Zarivach (b), J. Harms (a), A. Bashan (b), A. Tocilj (a), A. Yonath (a,b) and F. Franceschi (c), Nature, 413, 814-821 (2001); M. Pioletti (c), F. Schluenzen (a), J. Harms (a), R. Zarivach (b), M. Gluehmann (a), H. Avila (c), A. Bashan (a), H. Bartels (a), T. Auerbach (b), A. Yonath (a, b) and F. Franceschi (c) EMBO J. 20, 1829-1839 (2001).
(a) Max-Planck-Res. Unit for Ribosomal Structure, Hamburg (Germany)
(b) Dept. Structural Biology. Weizmann Inst. of Science, Rehovot (Israel)
(c) Max-Planck-Inst. for Molecular Genetics, Berlin (Germany)



