- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2001
- Life Sciences
- Sensory Rhodopsin Structure Provides First-Time Detailed Information of the Mechanism of Primitive Vision
Sensory Rhodopsin Structure Provides First-Time Detailed Information of the Mechanism of Primitive Vision
Sensory rhodopsin is a membrane protein that absorbs light and signals this information to flagellar components of the cell. The structure and function of this photoreceptor, which is related to rhodopsin in the mammalian retina, provides information on a primitive form of vision.
In studies on sensory rhodopsin, a team of researchers at UC Irvine, USA, led by H. Luecke, has used the very intense microfocus beamline, ID13, at the ESRF. They have elucidated a unique crystal structure [1] and for the first time are learning how this membrane protein changes shape to recognise the sun's emission spectrum. In addition, a surface binding site for downstream signalling proteins has been identified that does not exist in other members of the microbial rhodopsin family.
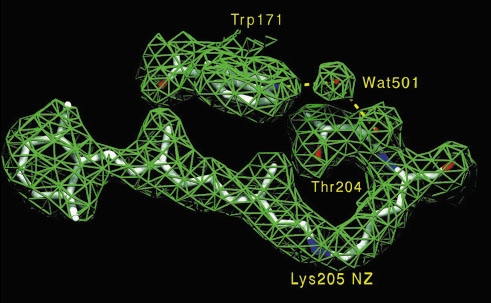 |
Fig. 22: Electron density map and corresponding molecular model of the retinal binding pocket. |
The three-dimensional structure of sensory rhodopsin II protein (NpSRII) (Figure 22) adds to our understanding of how it transforms when absorbing blue light, the most intense kind in the sun's emission spectrum. This highly specialised form of rhodopsin is a cell membrane protein found in a salt marsh-dwelling bacteria called Natronobacterium pharaonis. When sensory rhodopsin is activated, it sends a message through a second signaling protein, called a transducer, telling the cell how to react. A bacterium may want to avoid the harsh blue light and move toward a lower-energy form of light, when other forms of rhodopsin pump ions into the cell to store energy. The mechanism of spectral tuning is now being recognised and the structure also provides a crucial step to understanding the mechanisms involved with trans-membrane cell signalling. In capturing an image of sensory rhodopsin that absorbs blue light, a 1.1 Å shift of a charged group deep inside the molecule has been measured. This shift appears to be largely responsible for the change in the wavelength of absorption when compared with other rhodopsins.
With respect to a unique binding site for the signalling activity, inspection of the protein surface revealed an exposed amino acid, tyrosine, Tyr199, in the middle of the bilayer (Figure 23). This is believed to be one of the important sites where rhodopsin interacts with its transducer protein, enabling recognition of the very fundamental signal transduction with these molecules.
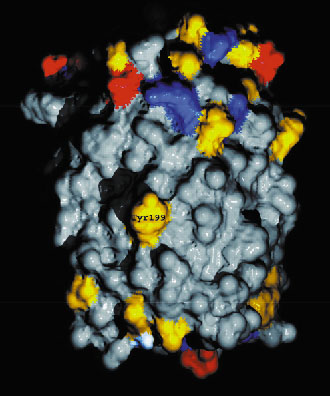 |
Fig. 23: Exposed conserved tyrosine in middle of bilayer. The NpSRII surface is coloured according to amino acid type (red, negatively charged; blue, positively charged; yellow, polar; grey, hydrophilic). |
The structures of membrane proteins, such as rhodopsin [2], are still relatively rare due to difficulties in isolation, purification and crystallisation. This new structural and functional information from rhodopsin is becoming increasingly important to understanding how G-protein coupled receptors (GPCRs) work. GPCRs, a superfamily of proteins that transduce signals across cell membranes, are proven to be excellent therapeutic targets. Nearly half of all known drugs act on GPCRs. While sensory rhodopsin is not a GPCR, it provides a model structure for study.
The team of UC Irvine plans to further explore crystal structures of rhodopsin molecules during different photocycle states in order to understand how the protein alters its shape to send messages.
References
[1] H. Luecke, B. Schobert, J.K. Lanyi, E.N. Spudich and J.L. Spudich, Science 293, 1499-1503 (2001).
[2] H. Luecke, B. Schobert, H.-T. Richter, J.-P. Cartailler and J.K. Lanyi, J. Mol. Biol. 291, 899-911 (1999).
Authors
H. Luecke (a), B. Schobert (a), J.K. Lanyi (a), E.N. Spudich (b) and J.L. Spudich (b).
(a) UCI's Department of Physiology and Biophysics, Irvine (USA)
(b) University of Texas Medical School, Houston (USA)



